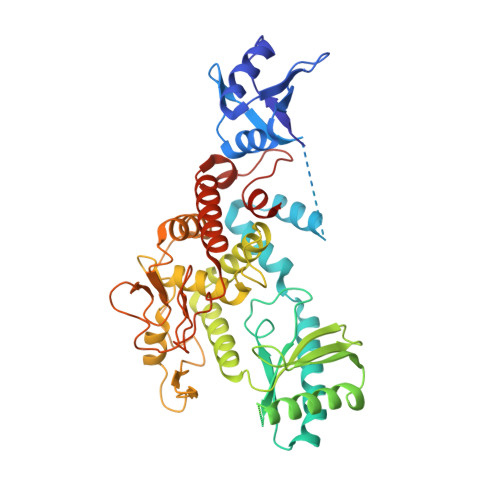
Crystal structures of U6 snRNA-specific terminal uridylyltransferase
Yamashita, S., Takagi, Y., Nagaike, T., Tomita, K.(2017) Nat Commun 8: 15788-15788
- PubMed: 28589955 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms15788
- Primary Citation Related Structures:
5WU1, 5WU2, 5WU3, 5WU4, 5WU5, 5WU6 - PubMed Abstract:
The terminal uridylyltransferase, TUT1, builds or repairs the 3'-oligo-uridylylated tail of U6 snRNA. The 3'-oligo-uridylylated tail is the Lsm-binding site for U4/U6 di-snRNP formation and U6 snRNA recycling for pre-mRNA splicing. Here, we report crystallographic and biochemical analyses of human TUT1, which revealed the mechanisms for the specific uridylylation of the 3'-end of U6 snRNA by TUT1. The O 2 and O 4 atoms of the UTP base form hydrogen bonds with the conserved His and Asn in the catalytic pocket, respectively, and TUT1 preferentially incorporates UMP onto the 3'-end of RNAs. TUT1 recognizes the entire U6 snRNA molecule by its catalytic domains, N-terminal RNA-recognition motifs and a previously unidentified C-terminal RNA-binding domain. Each domain recognizes specific regions within U6 snRNA, and the recognition is coupled with the domain movements and U6 snRNA structural changes. Hence, TUT1 functions as the U6 snRNA-specific terminal uridylyltransferase required for pre-mRNA splicing.
- Department of Computational Biology and Medical Sciences, Graduate School of Frontier Sciences, the University of Tokyo, Kashiwa, Chiba 277-8562, Japan.
Organizational Affiliation: