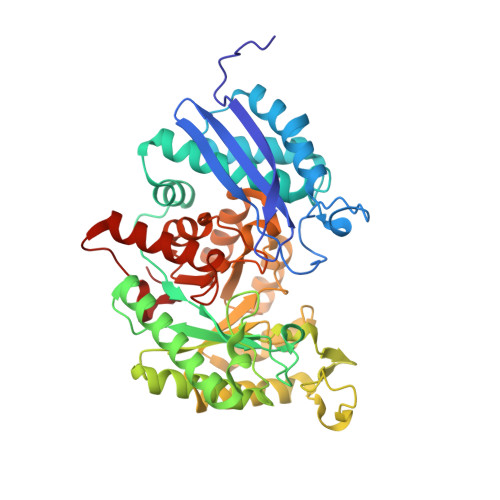
Crystal structure of enolase from Drosophila melanogaster.
Sun, C., Xu, B., Liu, X., Zhang, Z., Su, Z.(2017) Acta Crystallogr F Struct Biol Commun 73: 228-234
- PubMed: 28368282 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X17004022
- Primary Citation Related Structures:
5WRO - PubMed Abstract:
Enolase is an important enzyme in glycolysis and various biological processes. Its dysfunction is closely associated with diseases. Here, the enolase from Drosophila melanogaster (DmENO) was purified and crystallized. A crystal of DmENO diffracted to 2.0 Å resolution and belonged to space group R32. The structure was solved by molecular replacement. Like most enolases, DmENO forms a homodimer with conserved residues in the dimer interface. DmENO possesses an open conformation in this structure and contains conserved elements for catalytic activity. This work provides a structural basis for further functional and evolutionary studies of enolase.
- College of Chemical Engineering, Qingdao University of Science and Technology, Qingdao, Shandong 266042, People's Republic of China.
Organizational Affiliation: