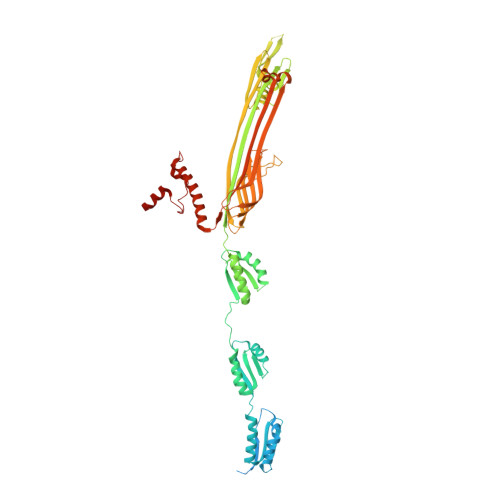
Structural insights into the secretin translocation channel in the type II secretion system
Yan, Z., Yin, M., Xu, D., Zhu, Y., Li, X.(2017) Nat Struct Mol Biol 24: 177-183
- PubMed: 28067918 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.3350
- Primary Citation Related Structures:
5WQ7, 5WQ8, 5WQ9 - PubMed Abstract:
The secretin GspD of the type II secretion system (T2SS) forms a channel across the outer membrane in Gram-negative bacteria to transport substrates from the periplasm to the extracellular milieu. The lack of an atomic-resolution structure of the GspD channel hinders the investigation of substrate translocation mechanism of T2SS. Here we report cryo-EM structures of two GspD channels (∼1 MDa), from Escherichia coli K12 and Vibrio cholerae, at ∼3 Å resolution. The structures reveal a pentadecameric channel architecture, wherein three rings of GspD N domains form the periplasmic channel. The secretin domain constitutes a novel double β-barrel channel, with at least 60 β-strands in each barrel, and is stabilized by S domains. The outer membrane channel is sealed by β-strand-enriched gates. On the basis of the partially open state captured, we proposed a detailed gate-opening mechanism. Our structures provide a structural basis for understanding the secretin superfamily and the mechanism of substrate translocation in T2SS.
- School of Life Sciences, Tsinghua University, Beijing, China.
Organizational Affiliation: