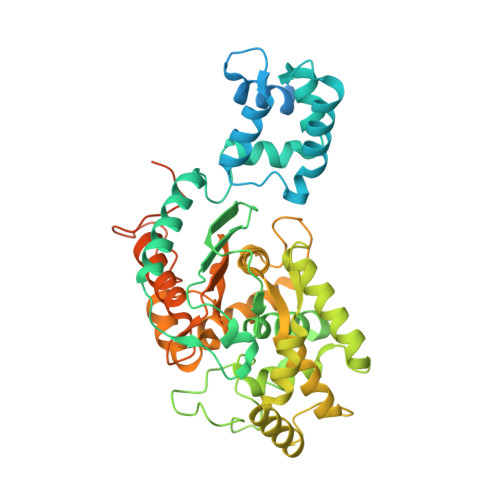
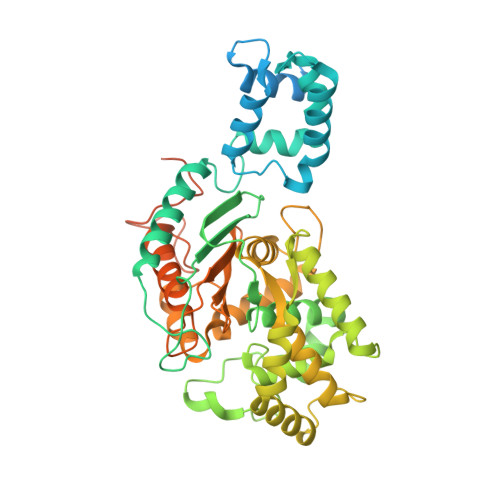
Characterization of the interactions of potent allosteric inhibitors with glutaminase C, a key enzyme in cancer cell glutamine metabolism.
Huang, Q., Stalnecker, C., Zhang, C., McDermott, L.A., Iyer, P., O'Neill, J., Reimer, S., Cerione, R.A., Katt, W.P.(2018) J Biological Chem 293: 3535-3545
- PubMed: 29317493 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M117.810101
- Primary Citation Related Structures:
5WJ6 - PubMed Abstract:
Altered glycolytic flux in cancer cells (the "Warburg effect") causes their proliferation to rely upon elevated glutamine metabolism ("glutamine addiction"). This requirement is met by the overexpression of glutaminase C (GAC), which catalyzes the first step in glutamine metabolism and therefore represents a potential therapeutic target. The small molecule CB-839 was reported to be more potent than other allosteric GAC inhibitors, including the parent compound bis-2-(5-phenylacetamido-1,2,4-thiadiazol-2-yl)ethyl (BPTES), and is in clinical trials. Recently, we described the synthesis of BPTES analogs having distinct saturated heterocyclic cores as a replacement for the flexible chain moiety, with improved microsomal stability relative to CB-839 and BPTES. Here, we show that one of these new compounds, UPGL00004, like CB-839, more potently inhibits the enzymatic activity of GAC, compared with BPTES. We also compare the abilities of UPGL00004, CB-839, and BPTES to directly bind to recombinant GAC and demonstrate that UPGL00004 has a similar binding affinity as CB-839 for GAC. We also show that UPGL00004 potently inhibits the growth of triple-negative breast cancer cells, as well as tumor growth when combined with the anti-vascular endothelial growth factor antibody bevacizumab. Finally, we compare the X-ray crystal structures for UPGL00004 and CB-839 bound to GAC, verifying that UPGL00004 occupies the same binding site as CB-839 or BPTES and that all three inhibitors regulate the enzymatic activity of GAC via a similar allosteric mechanism. These results provide insights regarding the potency of these inhibitors that will be useful in designing novel small-molecules that target a key enzyme in cancer cell metabolism.
- From the Cornell High Energy Synchrotron Source (CHESS) and.
Organizational Affiliation: