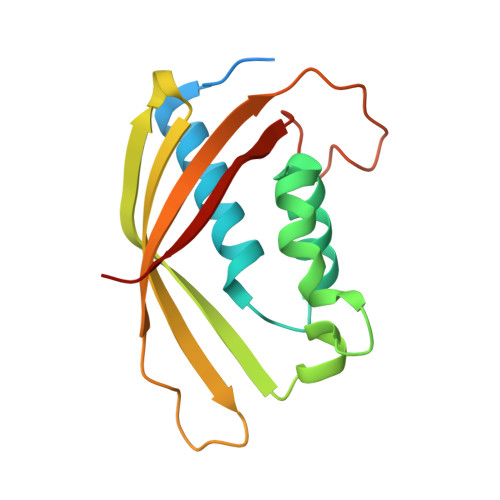
Fragment-based screening identifies novel targets for inhibitors of conjugative transfer of antimicrobial resistance by plasmid pKM101.
Casu, B., Arya, T., Bessette, B., Baron, C.(2017) Sci Rep 7: 14907-14907
- PubMed: 29097752 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-017-14953-1
- Primary Citation Related Structures:
5WIC, 5WII, 5WIO, 5WIP - PubMed Abstract:
The increasing frequency of antimicrobial resistance is a problem of global importance. Novel strategies are urgently needed to understand and inhibit antimicrobial resistance gene transmission that is mechanistically related to bacterial virulence functions. The conjugative transfer of plasmids by type IV secretion systems is a major contributor to antimicrobial resistance gene transfer. Here, we present a structure-based strategy to identify inhibitors of type IV secretion system-mediated bacterial conjugation. Using differential scanning fluorimetry we screened a fragment library and identified molecules that bind the essential TraE protein of the plasmid pKM101 conjugation machinery. Co-crystallization revealed that fragments bind two alternative sites of the protein and one of them is a novel inhibitor binding site. Based on the structural information on fragment binding we designed novel small molecules that have improved binding affinity. These molecules inhibit the dimerization of TraE, bind to both inhibitor binding sites on TraE and inhibit the conjugative transfer of plasmid pKM101. The strategy presented here is generally applicable for the structure-based design of inhibitors of antimicrobial resistance gene transfer and of bacterial virulence.
- Department of Biochemistry and Molecular Medicine, Faculty of Medicine, Université de Montréal, 2900 Boulevard Édouard-Montpetit, Montreal, QC, H3T 1J4, Canada.
Organizational Affiliation: