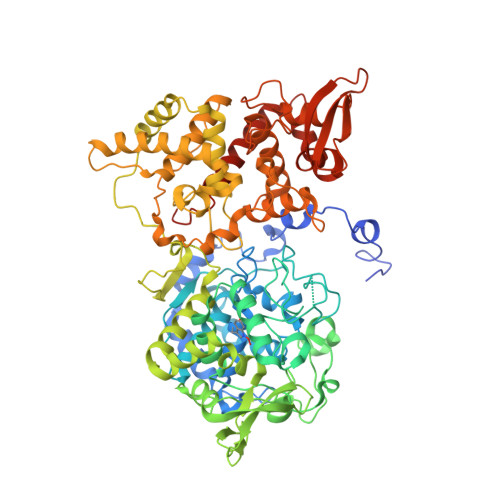
Structure, kinetics, molecular and redox properties of a cytosolic and developmentally regulated fungal catalase-peroxidase.
Vega-Garcia, V., Diaz-Vilchis, A., Saucedo-Vazquez, J.P., Solano-Peralta, A., Rudino-Pinera, E., Hansberg, W.(2018) Arch Biochem Biophys 640: 17-26
- PubMed: 29305053 Search on PubMed
- DOI: https://doi.org/10.1016/j.abb.2017.12.021
- Primary Citation Related Structures:
5WHQ, 5WHS - PubMed Abstract:
CAT-2, a cytosolic catalase-peroxidase (CP) from Neurospora crassa, which is induced during asexual spore formation, was heterologously expressed and characterized. CAT-2 had the Met-Tyr-Trp (M-Y-W) adduct required for catalase activity. Its K M for H 2 O 2 was micromolar for peroxidase and millimolar for catalase activity. A E m = -158 mV reduction potential value was obtained and the Soret band shift suggested a mixture of low and high spin ferric iron. CAT-2 EPR spectrum at 10 K indicated an axial and a rhombic component. With peroxyacetic acid (PAA), formation of Compound I* was observed with EPR. CAT-2 homodimer crystallographic structure contained two K + ions; Glu107 residues were displaced to bind them. CAT-2 showed the essential amino acid residues for activity in similar positions to other CPs. CAT-2 Arg426 is oriented towards the M-Y-W adduct, interacting with the deprotonated Tyr238 hydroxyl group. A perhydroxy modification of the indole nitrogen of Trp90 was oriented toward the catalytic His91. In contrast to cytochrome c peroxidase and ascorbate peroxidase, the catalase-peroxidase heme propionates are not exposed to the solvent. Together with other N. crassa enzymes that utilize H 2 O 2 as a substrate, CAT-2 has many tryptophan and proline residues at its surface, probably related to H 2 O 2 selection in water.
- Departamento de Biología Celular y del Desarrollo, Instituto de Fisiología Celular, Universidad Nacional Autónoma de México, UNAM, Mexico.
Organizational Affiliation: