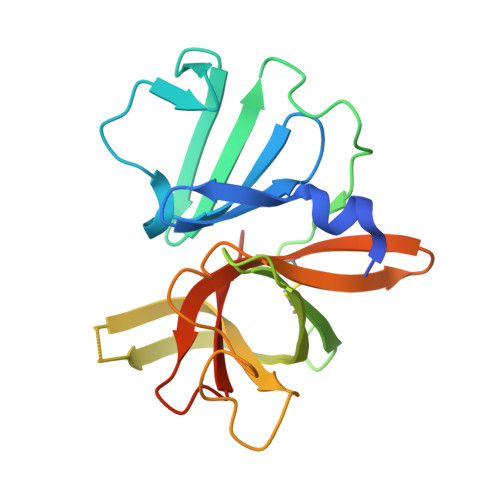
Structure-guided design, synthesis and evaluation of oxazolidinone-based inhibitors of norovirus 3CL protease.
Damalanka, V.C., Kim, Y., Galasiti Kankanamalage, A.C., Rathnayake, A.D., Mehzabeen, N., Battaile, K.P., Lovell, S., Nguyen, H.N., Lushington, G.H., Chang, K.O., Groutas, W.C.(2017) Eur J Med Chem 143: 881-890
- PubMed: 29227928 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.ejmech.2017.12.014
- Primary Citation Related Structures:
5WEJ - PubMed Abstract:
Acute nonbacterial gastroenteritis caused by noroviruses constitutes a global public health concern and a significant economic burden. There are currently no small molecule therapeutics or vaccines for the treatment of norovirus infections. A structure-guided approach was utilized in the design of a series of inhibitors of norovirus 3CL protease that embody an oxazolidinone ring as a novel design element for attaining optimal binding interactions. Low micromolar cell-permeable inhibitors that display anti-norovirus activity have been identified. The mechanism of action, mode of binding, and structural rearrangements associated with the interaction of the inhibitors and the enzyme were elucidated using X-ray crystallography.
- Department of Chemistry, Wichita State University, Wichita, KS 67260, USA.
Organizational Affiliation: