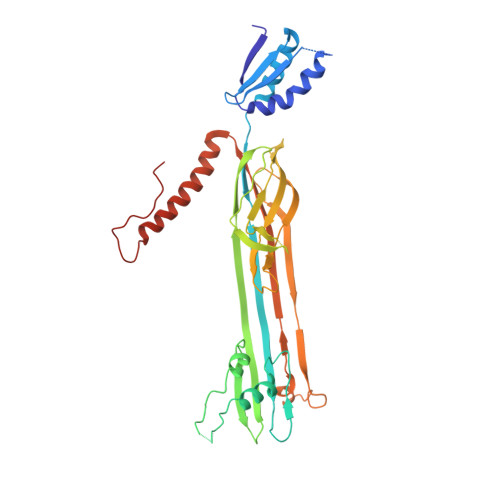
Structure and Membrane Topography of the Vibrio-Type Secretin Complex from the Type 2 Secretion System of Enteropathogenic Escherichia coli.
Hay, I.D., Belousoff, M.J., Dunstan, R.A., Bamert, R.S., Lithgow, T.(2018) J Bacteriol 200
- PubMed: 29084860 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.00521-17
- Primary Citation Related Structures:
5W68 - PubMed Abstract:
The β-barrel assembly machinery (BAM) complex is the core machinery for the assembly of β-barrel membrane proteins, and inhibition of BAM complex activity is lethal to bacteria. Discovery of integral membrane proteins that are key to pathogenesis and yet do not require assistance from the BAM complex raises the question of how these proteins assemble into bacterial outer membranes. Here, we address this question through a structural analysis of the type 2 secretion system (T2SS) secretin from enteropathogenic Escherichia coli O127:H6 strain E2348/69. Long β-strands assemble into a barrel extending 17 Å through and beyond the outer membrane, adding insight to how these extensive β-strands are assembled into the E. coli outer membrane. The substrate docking chamber of this secretin is shown to be sufficient to accommodate the substrate mucinase SteC. IMPORTANCE In order to cause disease, bacterial pathogens inhibit immune responses and induce pathology that will favor their replication and dissemination. In Gram-negative bacteria, these key attributes of pathogenesis depend on structures assembled into or onto the outer membrane. One of these is the T2SS. The Vibrio -type T2SS mediates cholera toxin secretion in Vibrio cholerae , and in Escherichia coli O127:H6 strain E2348/69, the same machinery mediates secretion of the mucinases that enable the pathogen to penetrate intestinal mucus and thereby establish deadly infections.
- Infection and Immunity Program, Biomedicine Discovery Institute and Department of Microbiology, Monash University, Clayton, Australia.
Organizational Affiliation: