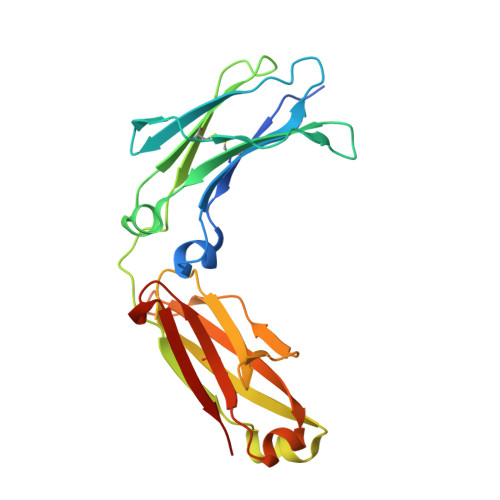
Structural characterization of the Man5 glycoform of human IgG3 Fc.
Shah, I.S., Lovell, S., Mehzabeen, N., Battaile, K.P., Tolbert, T.J.(2017) Mol Immunol 92: 28-37
- PubMed: 29031045 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molimm.2017.10.001
- Primary Citation Related Structures:
5W38 - PubMed Abstract:
Immunoglobulin G (IgG) consists of four subclasses in humans: IgG1, IgG2, IgG3 and IgG4, which are highly conserved but have unique differences that result in subclass-specific effector functions. Though IgG1 is the most extensively studied IgG subclass, study of other subclasses is important to understand overall immune function and for development of new therapeutics. When compared to IgG1, IgG3 exhibits a similar binding profile to Fcγ receptors and stronger activation of complement. All IgG subclasses are glycosylated at N297, which is required for Fcγ receptor and C1q complement binding as well as maintaining optimal Fc conformation. We have determined the crystal structure of homogenously glycosylated human IgG3 Fc with a GlcNAc 2 Man 5 (Man5) high mannose glycoform at 1.8Å resolution and compared its structural features with published structures from the other IgG subclasses. Although the overall structure of IgG3 Fc is similar to that of other subclasses, some structural perturbations based on sequence differences were revealed. For instance, the presence of R435 in IgG3 (and H435 in the other IgG subclasses) has been implicated to result in IgG3-specific properties related to binding to protein A, protein G and the neonatal Fc receptor (FcRn). The IgG3 Fc structure helps to explain some of these differences. Additionally, protein-glycan contacts observed in the crystal structure appear to correlate with IgG3 affinity for Fcγ receptors as shown by binding studies with IgG3 Fc glycoforms. Finally, this IgG3 Fc structure provides a template for further studies aimed at engineering the Fc for specific gain of function.
- Department of Pharmaceutical Chemistry, University of Kansas, Lawrence, KS, USA.
Organizational Affiliation: