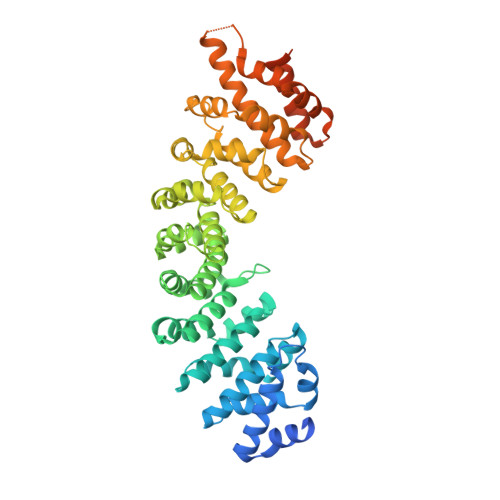
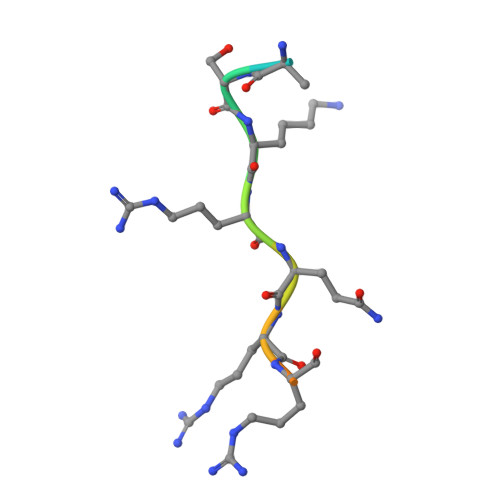
Nuclear transport of the Neurospora crassa NIT-2 transcription factor is mediated by importin-alpha.
Bernardes, N.E., Takeda, A.A.S., Dreyer, T.R., Cupertino, F.B., Virgilio, S., Pante, N., Bertolini, M.C., Fontes, M.R.M.(2017) Biochem J 474: 4091-4104
- PubMed: 29054975 Search on PubMed
- DOI: https://doi.org/10.1042/BCJ20170654
- Primary Citation Related Structures:
5VQI - PubMed Abstract:
The Neurospora crassa NIT-2 transcription factor belongs to the GATA transcription factor family and plays a fundamental role in the regulation of nitrogen metabolism. Because NIT-2 acts by accessing DNA inside the nucleus, understanding the nuclear import process of NIT-2 is necessary to characterize its function. Thus, in the present study, NIT-2 nuclear transport was investigated using a combination of biochemical, cellular, and biophysical methods. A complemented strain that produced an sfGFP-NIT-2 fusion protein was constructed, and nuclear localization assessments were made under conditions that favored protein translocation to the nucleus. Nuclear translocation was also investigated using HeLa cells, which showed that the putative NIT-2 nuclear localization sequence (NLS; 915 TISSKRQRRHSKS 927 ) was recognized by importin-α and that subsequent transport occurred via the classical import pathway. The interaction between the N. crassa importin-α (NcImpα) and the NIT-2 NLS was quantified with calorimetric assays, leading to the observation that the peptide bound to two sites with different affinities, which is typical of a monopartite NLS sequence. The crystal structure of the NcImpα/NIT-2 NLS complex was solved and revealed that the NIT-2 peptide binds to NcImpα with the major NLS-binding site playing a primary role. This result contrasts other recent studies that suggested a major role for the minor NLS-binding site in importin-α from the α2 family, indicating that both sites can be used for different cargo proteins according to specific metabolic requirements.
- Departamento de Física e Biofísica, Instituto de Biociências, Universidade Estadual Paulista (UNESP), Botucatu, São Paulo, Brazil.
Organizational Affiliation: