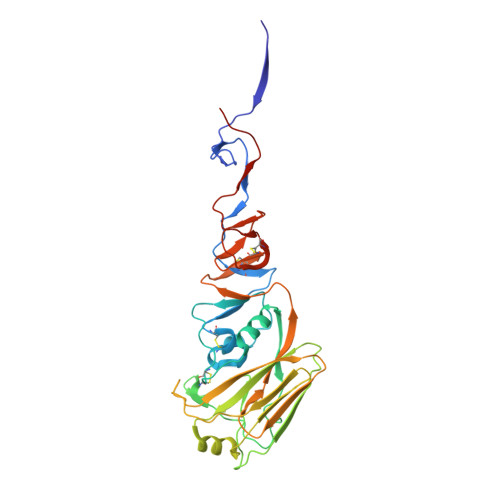
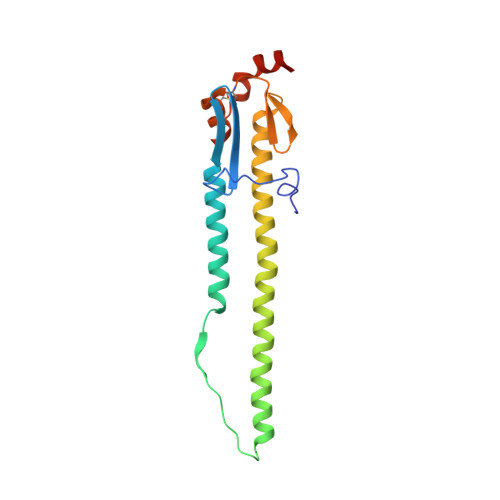
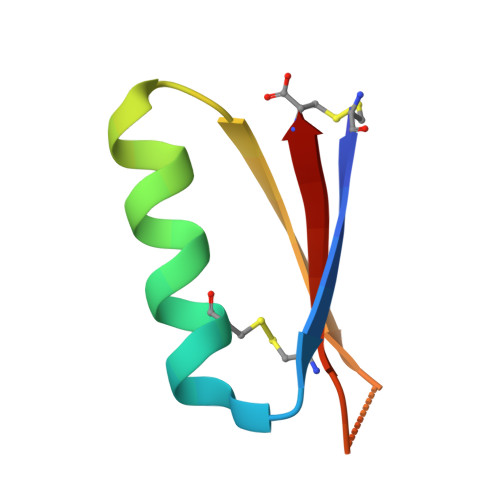
Massively parallel de novo protein design for targeted therapeutics.
Chevalier, A., Silva, D.A., Rocklin, G.J., Hicks, D.R., Vergara, R., Murapa, P., Bernard, S.M., Zhang, L., Lam, K.H., Yao, G., Bahl, C.D., Miyashita, S.I., Goreshnik, I., Fuller, J.T., Koday, M.T., Jenkins, C.M., Colvin, T., Carter, L., Bohn, A., Bryan, C.M., Fernandez-Velasco, D.A., Stewart, L., Dong, M., Huang, X., Jin, R., Wilson, I.A., Fuller, D.H., Baker, D.(2017) Nature 550: 74-79
- PubMed: 28953867 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature23912
- Primary Citation Related Structures:
5VID, 5VLI, 5VMR - PubMed Abstract:
De novo protein design holds promise for creating small stable proteins with shapes customized to bind therapeutic targets. We describe a massively parallel approach for designing, manufacturing and screening mini-protein binders, integrating large-scale computational design, oligonucleotide synthesis, yeast display screening and next-generation sequencing. We designed and tested 22,660 mini-proteins of 37-43 residues that target influenza haemagglutinin and botulinum neurotoxin B, along with 6,286 control sequences to probe contributions to folding and binding, and identified 2,618 high-affinity binders. Comparison of the binding and non-binding design sets, which are two orders of magnitude larger than any previously investigated, enabled the evaluation and improvement of the computational model. Biophysical characterization of a subset of the binder designs showed that they are extremely stable and, unlike antibodies, do not lose activity after exposure to high temperatures. The designs elicit little or no immune response and provide potent prophylactic and therapeutic protection against influenza, even after extensive repeated dosing.
- Department of Biochemistry, University of Washington, Seattle, Washington 98195, USA.
Organizational Affiliation: