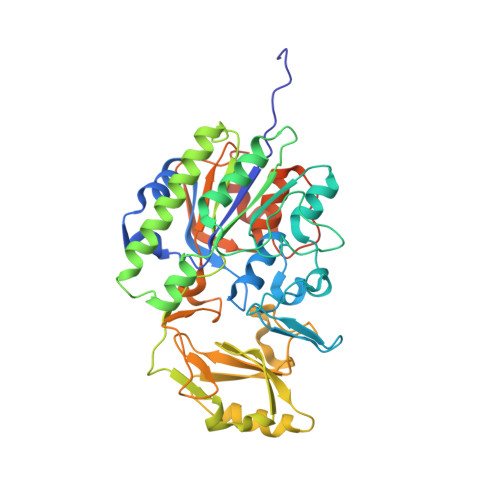
A key tyrosine substitution restricts nucleotide hydrolysis by the ectoenzyme NPP5.
Gorelik, A., Randriamihaja, A., Illes, K., Nagar, B.(2017) FEBS J 284: 3718-3726
- PubMed: 28898552 Search on PubMed
- DOI: https://doi.org/10.1111/febs.14266
- Primary Citation Related Structures:
5VEM, 5VEN, 5VEO - PubMed Abstract:
The ecto-nucleotide pyrophosphatase/phosphodiesterase (NPP) family of proteins mediates purinergic signaling by degrading extracellular nucleotides and also participates in phospholipid metabolism. NPP5 (ENPP5) is the least characterized member of this group and its specific role is unknown. This enzyme does not display activity on certain nucleotides and on other typical NPP substrates. In order to gain insights into its function, we determined the crystal structure of human and murine NPP5. Structural comparison with close homologs revealed a key phenylalanine to tyrosine substitution that prevents efficient hydrolysis of nucleotide diphosphates and triphosphates; reversal of this mutation enabled degradation of these molecules. Interestingly, NPP5 is able to cleave nicotinamide adenine dinucleotide (NAD), suggesting a potential role of this enzyme in NAD-based neurotransmission. An NPP5-specific metal binding motif is found adjacent to the active site, although its significance is unclear. These findings expand our understanding of substrate specificity within the NPP family. Structural data are available in the Protein Data Bank under the accession numbers 5VEM, 5VEN, and 5VEO.
- Department of Biochemistry and Groupe de Recherche Axé sur la Structure des Protéines, McGill University, Montreal, Canada.
Organizational Affiliation: