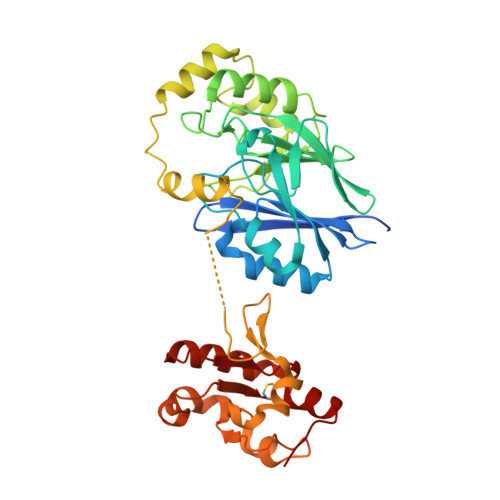
Structural and biochemical analyses indicate that a bacterial persulfide dioxygenase-rhodanese fusion protein functions in sulfur assimilation.
Motl, N., Skiba, M.A., Kabil, O., Smith, J.L., Banerjee, R.(2017) J Biological Chem 292: 14026-14038
- PubMed: 28684420
- DOI: https://doi.org/10.1074/jbc.M117.790170
- Primary Citation Related Structures:
5VE3, 5VE4, 5VE5 - PubMed Abstract:
Hydrogen sulfide (H 2 S) is a signaling molecule that is toxic at elevated concentrations. In eukaryotes, it is cleared via a mitochondrial sulfide oxidation pathway, which comprises sulfide quinone oxidoreductase, persulfide dioxygenase (PDO), rhodanese, and sulfite oxidase and converts H 2 S to thiosulfate and sulfate. Natural fusions between the non-heme iron containing PDO and rhodanese, a thiol sulfurtransferase, exist in some bacteria. However, little is known about the role of the PDO-rhodanese fusion (PRF) proteins in sulfur metabolism. Herein, we report the kinetic properties and the crystal structure of a PRF from the Gram-negative endophytic bacterium Burkholderia phytofirmans The crystal structures of wild-type PRF and a sulfurtransferase-inactivated C314S mutant with and without glutathione were determined at 1.8, 2.4, and 2.7 Å resolution, respectively. We found that the two active sites are distant and do not show evidence of direct communication. The B. phytofirmans PRF exhibited robust PDO activity and preferentially catalyzed sulfur transfer in the direction of thiosulfate to sulfite and glutathione persulfide; sulfur transfer in the reverse direction was detectable only under limited turnover conditions. Together with the kinetic data, our bioinformatics analysis reveals that B. phytofirmans PRF is poised to metabolize thiosulfate to sulfite in a sulfur assimilation pathway rather than in sulfide stress response as seen, for example, with the Staphylococcus aureus PRF or sulfide oxidation and disposal as observed with the homologous mammalian proteins.
- From the Department of Biological Chemistry, University of Michigan Medical Center, Ann Arbor, Michigan 48109-0600.
Organizational Affiliation: