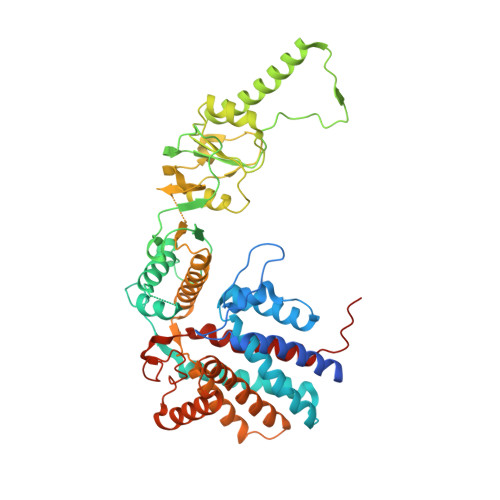
Structure of the human TRiC/CCT Subunit 5 associated with hereditary sensory neuropathy.
Pereira, J.H., McAndrew, R.P., Sergeeva, O.A., Ralston, C.Y., King, J.A., Adams, P.D.(2017) Sci Rep 7: 3673-3673
- PubMed: 28623285 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-017-03825-3
- Primary Citation Related Structures:
5UYX, 5UYZ - PubMed Abstract:
The human chaperonin TRiC consists of eight non-identical subunits, and its protein-folding activity is critical for cellular health. Misfolded proteins are associated with many human diseases, such as amyloid diseases, cancer, and neuropathies, making TRiC a potential therapeutic target. A detailed structural understanding of its ATP-dependent folding mechanism and substrate recognition is therefore of great importance. Of particular health-related interest is the mutation Histidine 147 to Arginine (H147R) in human TRiC subunit 5 (CCT5), which has been associated with hereditary sensory neuropathy. In this paper, we describe the crystal structures of CCT5 and the CCT5-H147R mutant, which provide important structural information for this vital protein-folding machine in humans. This first X-ray crystallographic study of a single human CCT subunit in the context of a hexadecameric complex can be expanded in the future to the other 7 subunits that form the TRiC complex.
- Molecular Biophysics and Integrated Bioimaging Division, Lawrence Berkeley National Laboratory, Berkeley, CA, 94720, USA.
Organizational Affiliation: