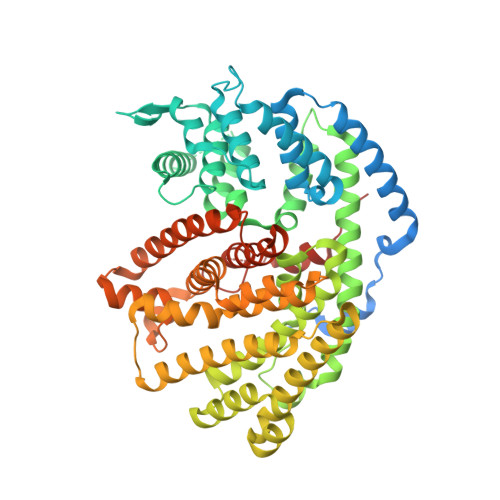
Functional and Structural Characterization of a (+)-Limonene Synthase from Citrus sinensis.
Morehouse, B.R., Kumar, R.P., Matos, J.O., Olsen, S.N., Entova, S., Oprian, D.D.(2017) Biochemistry 56: 1706-1715
- PubMed: 28272875 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.7b00143
- Primary Citation Related Structures:
5UV0 - PubMed Abstract:
Terpenes make up the largest and most diverse class of natural compounds and have important commercial and medical applications. Limonene is a cyclic monoterpene (C 10 ) present in nature as two enantiomers, (+) and (-), which are produced by different enzymes. The mechanism of production of the (-)-enantiomer has been studied in great detail, but to understand how enantiomeric selectivity is achieved in this class of enzymes, it is important to develop a thorough biochemical description of enzymes that generate (+)-limonene, as well. Here we report the first cloning and biochemical characterization of a (+)-limonene synthase from navel orange (Citrus sinensis). The enzyme obeys classical Michaelis-Menten kinetics and produces exclusively the (+)-enantiomer. We have determined the crystal structure of the apoprotein in an "open" conformation at 2.3 Å resolution. Comparison with the structure of (-)-limonene synthase (Mentha spicata), which is representative of a fully closed conformation (Protein Data Bank entry 2ONG ), reveals that the short H-α1 helix moves nearly 5 Å inward upon substrate binding, and a conserved Tyr flips to point its hydroxyl group into the active site.
- Department of Biochemistry, Brandeis University , Waltham, Massachusetts 02454, United States.
Organizational Affiliation: