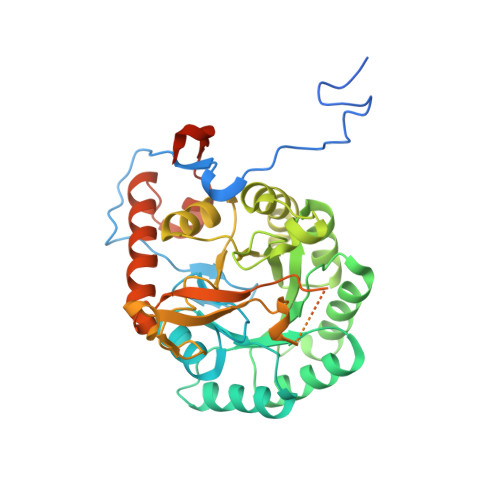
Crystal Structure of the Catalytic Domain of the Inosine Monophosphate Dehydrogenase from Mycobacterium tuberculosis in the presence of TBK6
Kim, Y., Makowska-Grzyska, M., Maltseva, N., Mulligan, R., Gu, M., Sacchettini, J., Anderson, W.F., Joachimiak, A., Center for Structural Genomics of Infectious Diseases (CSGID)To be published.