Crystal structure of the F255A mutant Kir3.1 cytoplasmic pore domain
Meinke, G., Bohm, A., Noujaim, S.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| G protein-activated inward rectifier potassium channel 1 | 210 | Anolis carolinensis, Mus musculus This entity is chimeric | Mutation(s): 1 Gene Names: LOC100559481, Kcnj3, Girk1 | 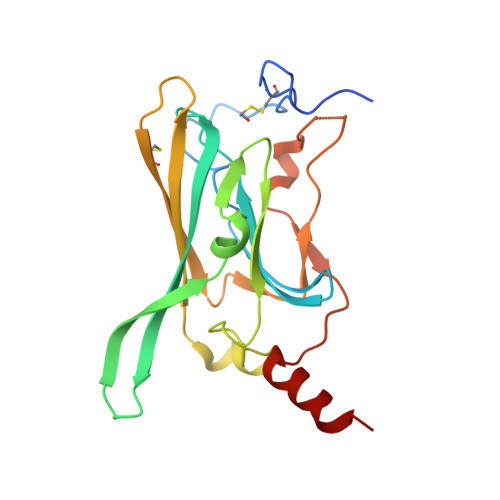 | |
UniProt & NIH Common Fund Data Resources | |||||
IMPC: MGI:104742 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Groups | H9GIV0P63250 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| CME Query on CME | A | L-PEPTIDE LINKING | C5 H11 N O3 S2 |  | CYS |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 80.07 | α = 90 |
| b = 80.07 | β = 90 |
| c = 85.04 | γ = 90 |
| Software Name | Purpose |
|---|---|
| SCALA | data scaling |
| PHASER | phasing |
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| MOSFLM | data reduction |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Heart, Lung, and Blood Institute (NIH/NHLBI) | United States | -- |