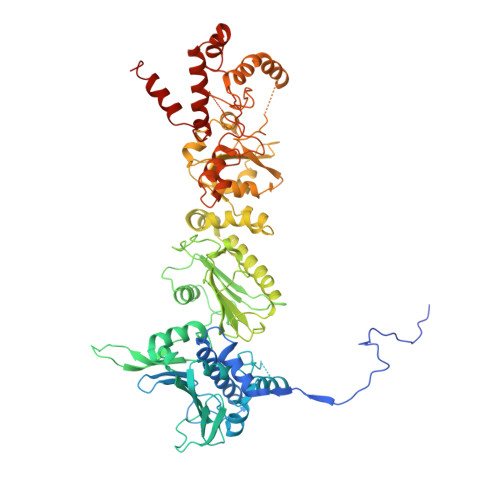
Structural and Functional Analysis of GRP94 in the Closed State Reveals an Essential Role for the Pre-N Domain and a Potential Client-Binding Site.
Huck, J.D., Que, N.L., Hong, F., Li, Z., Gewirth, D.T.(2017) Cell Rep 20: 2800-2809
- PubMed: 28930677 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2017.08.079
- Primary Citation Related Structures:
5ULS - PubMed Abstract:
Hsp90 chaperones undergo ATP-driven conformational changes during the maturation of client proteins, populating a closed state upon ATP binding in which the N-terminal domains of the homodimer form a second inter-protomer dimer interface. A structure of GRP94, the endoplasmic reticulum hsp90, in a closed conformation has not been described, and the determinants that regulate closure are not well understood. Here, we determined the 2.6-Å structure of AMPPNP-bound GRP94 in the closed dimer conformation. The structure includes the pre-N domain, a region preceding the N-terminal domain that is highly conserved in GRP94, but not in other hsp90s. We show that the GRP94 pre-N domain is essential for client maturation, and we identify the pre-N domain as an important regulator of ATPase rates and dimer closure. The structure also reveals a GRP94:polypeptide interaction that partially mimics a client-bound state. The results provide structural insight into the ATP-dependent client maturation process of GRP94.
- Hauptman-Woodward Medical Research Institute, Buffalo, NY, USA; Department of Structural Biology, University at Buffalo, Buffalo, NY, USA.
Organizational Affiliation: