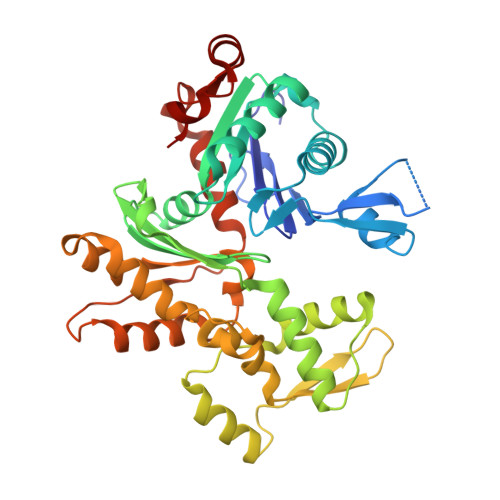
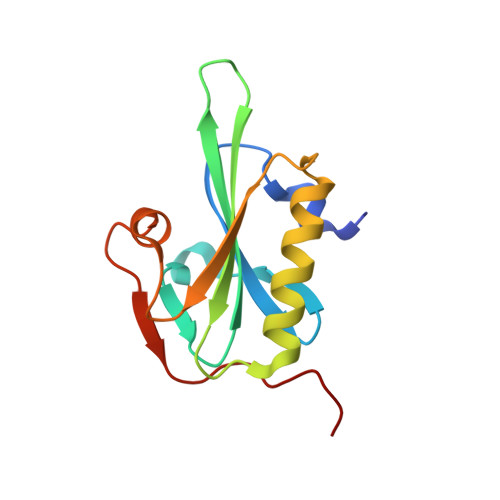
Catastrophic disassembly of actin filaments via Mical-mediated oxidation.
Grintsevich, E.E., Ge, P., Sawaya, M.R., Yesilyurt, H.G., Terman, J.R., Zhou, Z.H., Reisler, E.(2017) Nat Commun 8: 2183-2183
- PubMed: 29259197 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-02357-8
- Primary Citation Related Structures:
5UBO, 6AV9, 6AVB - PubMed Abstract:
Actin filament assembly and disassembly are vital for cell functions. MICAL Redox enzymes are important post-translational effectors of actin that stereo-specifically oxidize actin's M44 and M47 residues to induce cellular F-actin disassembly. Here we show that Mical-oxidized (Mox) actin can undergo extremely fast (84 subunits/s) disassembly, which depends on F-actin's nucleotide-bound state. Using near-atomic resolution cryoEM reconstruction and single filament TIRF microscopy we identify two dynamic and structural states of Mox-actin. Modeling actin's D-loop region based on our 3.9 Å cryoEM reconstruction suggests that oxidation by Mical reorients the side chain of M44 and induces a new intermolecular interaction of actin residue M47 (M47-O-T351). Site-directed mutagenesis reveals that this interaction promotes Mox-actin instability. Moreover, we find that Mical oxidation of actin allows for cofilin-mediated severing even in the presence of inorganic phosphate. Thus, in conjunction with cofilin, Mical oxidation of actin promotes F-actin disassembly independent of the nucleotide-bound state.
- Department of Chemistry and Biochemistry, University of California (UCLA), Los Angeles, CA, 90095, USA.
Organizational Affiliation: