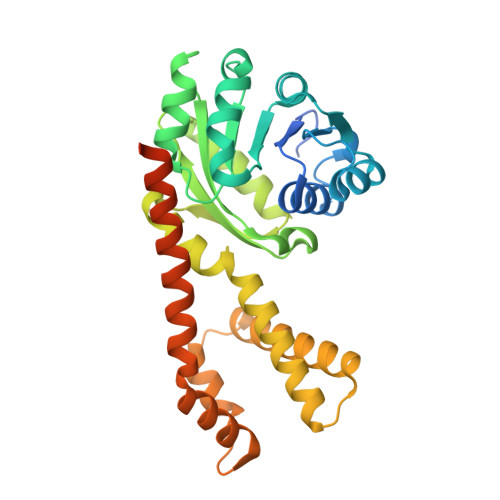
Resolving the cofactor-binding site in the proline biosynthetic enzyme human pyrroline-5-carboxylate reductase 1.
Christensen, E.M., Patel, S.M., Korasick, D.A., Campbell, A.C., Krause, K.L., Becker, D.F., Tanner, J.J.(2017) J Biological Chem 292: 7233-7243
- PubMed: 28258219 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M117.780288
- Primary Citation Related Structures:
5UAT, 5UAU, 5UAV, 5UAW, 5UAX - PubMed Abstract:
Pyrroline-5-carboxylate reductase (PYCR) is the final enzyme in proline biosynthesis, catalyzing the NAD(P)H-dependent reduction of Δ 1 -pyrroline-5-carboxylate (P5C) to proline. Mutations in the PYCR1 gene alter mitochondrial function and cause the connective tissue disorder cutis laxa. Furthermore, PYCR1 is overexpressed in multiple cancers, and the PYCR1 knock-out suppresses tumorigenic growth, suggesting that PYCR1 is a potential cancer target. However, inhibitor development has been stymied by limited mechanistic details for the enzyme, particularly in light of a previous crystallographic study that placed the cofactor-binding site in the C-terminal domain rather than the anticipated Rossmann fold of the N-terminal domain. To fill this gap, we report crystallographic, sedimentation-velocity, and kinetics data for human PYCR1. Structures of binary complexes of PYCR1 with NADPH or proline determined at 1.9 Å resolution provide insight into cofactor and substrate recognition. We see NADPH bound to the Rossmann fold, over 25 Å from the previously proposed site. The 1.85 Å resolution structure of a ternary complex containing NADPH and a P5C/proline analog provides a model of the Michaelis complex formed during hydride transfer. Sedimentation velocity shows that PYCR1 forms a concentration-dependent decamer in solution, consistent with the pentamer-of-dimers assembly seen crystallographically. Kinetic and mutational analysis confirmed several features seen in the crystal structure, including the importance of a hydrogen bond between Thr-238 and the substrate as well as limited cofactor discrimination.
- From the Departments of Chemistry and.
Organizational Affiliation: