New Ulvan-Degrading Polysaccharide Lyase Family: Structure and Catalytic Mechanism Suggests Convergent Evolution of Active Site Architecture.
Ulaganathan, T., Boniecki, M.T., Foran, E., Buravenkov, V., Mizrachi, N., Banin, E., Helbert, W., Cygler, M.(2017) ACS Chem Biol 12: 1269-1280
- PubMed: 28290654 Search on PubMed
- DOI: https://doi.org/10.1021/acschembio.7b00126
- Primary Citation Related Structures:
5UAM, 5UAS - PubMed Abstract:
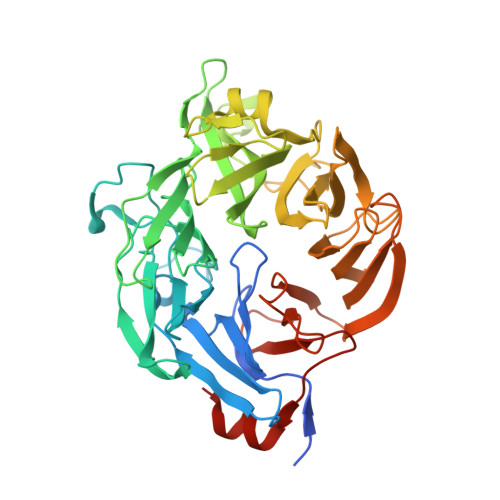
Ulvan is a complex sulfated polysaccharide biosynthesized by green seaweed and contains predominantly rhamnose, xylose, and uronic acid sugars. Ulvan-degrading enzymes have only recently been identified and added to the CAZy ( www.cazy.org ) database as family PL24, but neither their structure nor catalytic mechanism(s) are yet known. Several homologous, new ulvan lyases, have been discovered in Pseudoalteromonas sp. strain PLSV, Alteromonas LOR, and Nonlabens ulvanivorans, defining a new family PL25, with the lyase encoded by the gene PLSV_3936 being one of them. This enzyme cleaves the glycosidic bond between 3-sulfated rhamnose (R3S) and glucuronic acid (GlcA) or iduronic acid (IdoA) via a β-elimination mechanism. We report the crystal structure of PLSV_3936 and its complex with a tetrasaccharide substrate. PLSV_3936 folds into a seven-bladed β-propeller, with each blade consisting of four antiparallel β-strands. Sequence conservation analysis identified a highly conserved region lining at one end of a deep crevice on the protein surface. The putative active site was identified by mutagenesis and activity measurements. Crystal structure of the enzyme with a bound tetrasaccharide substrate confirmed the identity of base and acid residues and allowed determination of the catalytic mechanism and also the identification of residues neutralizing the uronic acid carboxylic group. The PLSV_3936 structure provides an example of a convergent evolution among polysaccharide lyases toward a common active site architecture embedded in distinct folds.
- Department of Biochemistry, University of Saskatchewan , Saskatoon, Saskatchewan S7N 5E5, Canada.
Organizational Affiliation: