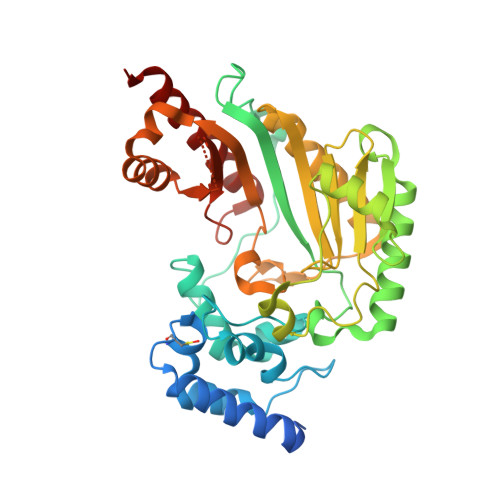
Biochemical and structural characterization of a novel arginine kinase from the spider Polybetes pythagoricus.
Laino, A., Lopez-Zavala, A.A., Garcia-Orozco, K.D., Carrasco-Miranda, J.S., Santana, M., Stojanoff, V., Sotelo-Mundo, R.R., Garcia, C.F.(2017) PeerJ 5: e3787-e3787
- PubMed: 28924503 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7717/peerj.3787
- Primary Citation Related Structures:
5U8E, 5U92 - PubMed Abstract:
Energy buffering systems are key for homeostasis during variations in energy supply. Spiders are the most important predators for insects and therefore key in terrestrial ecosystems. From biomedical interest, spiders are important for their venoms and as a source of potent allergens, such as arginine kinase (AK, EC 2.7.3.3). AK is an enzyme crucial for energy metabolism, keeping the pool of phosphagens in invertebrates, and also an allergen for humans. In this work, we studied AK from the Argentininan spider Polybetes pythagoricus ( Pp AK), from its complementary DNA to the crystal structure. The Pp AK cDNA from muscle was cloned, and it is comprised of 1068 nucleotides that encode a 384-amino acids protein, similar to other invertebrate AKs. The apparent Michaelis-Menten kinetic constant ( K m ) was 1.7 mM with a k cat of 75 s -1 . Two crystal structures are presented, the apo Pv AK and Pp AK bound to arginine, both in the open conformation with the active site lid (residues 310-320) completely disordered. The guanidino group binding site in the apo structure appears to be organized to accept the arginine substrate. Finally, these results contribute to knowledge of mechanistic details of the function of arginine kinase.
- Instituto de Investigaciones Bioquímicas de La Plata "Dr. Prof. Rodolfo R. Brenner", Universidad Nacional de La Plata, La Plata, Buenos Aires, Argentina.
Organizational Affiliation: