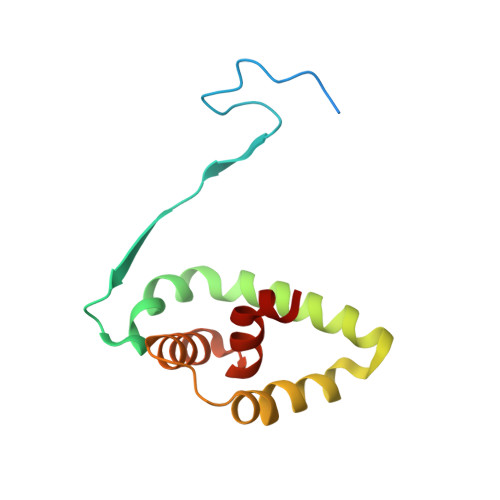
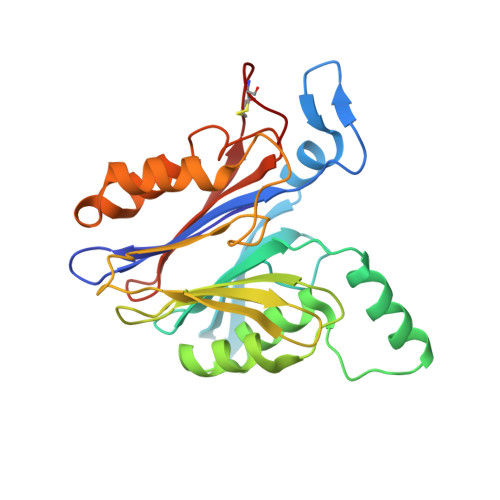
Structural basis for the activation of acid ceramidase.
Gebai, A., Gorelik, A., Li, Z., Illes, K., Nagar, B.(2018) Nat Commun 9: 1621-1621
- PubMed: 29692406 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-03844-2
- Primary Citation Related Structures:
5U7Z, 5U81, 5U84 - PubMed Abstract:
Acid ceramidase (aCDase, ASAH1) hydrolyzes lysosomal membrane ceramide into sphingosine, the backbone of all sphingolipids, to regulate many cellular processes. Abnormal function of aCDase leads to Farber disease, spinal muscular atrophy with progressive myoclonic epilepsy, and is associated with Alzheimer's, diabetes, and cancer. Here, we present crystal structures of mammalian aCDases in both proenzyme and autocleaved forms. In the proenzyme, the catalytic center is buried and protected from solvent. Autocleavage triggers a conformational change exposing a hydrophobic channel leading to the active site. Substrate modeling suggests distinct catalytic mechanisms for substrate hydrolysis versus autocleavage. A hydrophobic surface surrounding the substrate binding channel appears to be a site of membrane attachment where the enzyme accepts substrates facilitated by the accessory protein, saposin-D. Structural mapping of disease mutations reveals that most would destabilize the protein fold. These results will inform the rational design of aCDase inhibitors and recombinant aCDase for disease therapeutics.
- Department of Biochemistry and Groupe de Recherche Axé sur la Structure des Protéines, McGill University, Montreal, QC, H3G 0B1, Canada.
Organizational Affiliation: