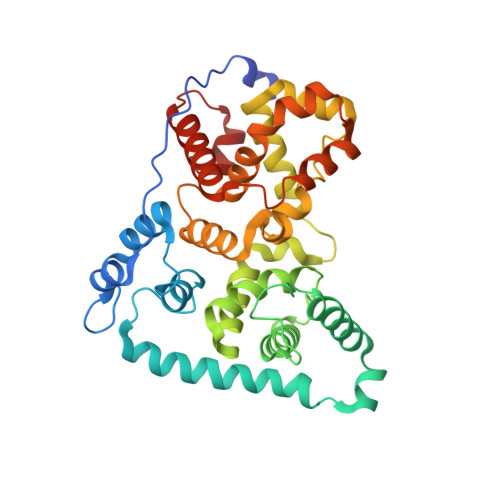
Crystal structure of TBC1D15 GTPase-activating protein (GAP) domain and its activity on Rab GTPases.
Chen, Y.N., Gu, X., Zhou, X.E., Wang, W., Cheng, D., Ge, Y., Ye, F., Xu, H.E., Lv, Z.(2017) Protein Sci 26: 834-846
- PubMed: 28168758
- DOI: https://doi.org/10.1002/pro.3132
- Primary Citation of Related Structures:
5TUB, 5TUC - PubMed Abstract:
TBC1D15 belongs to the TBC (Tre-2/Bub2/Cdc16) domain family and functions as a GTPase-activating protein (GAP) for Rab GTPases. So far, the structure of TBC1D15 or the TBC1D15·Rab complex has not been determined, thus, its catalytic mechanism on Rab GTPases is still unclear. In this study, we solved the crystal structures of the Shark and Sus TBC1D15 GAP domains, to 2.8 Å and 2.5 Å resolution, respectively. Shark-TBC1D15 and Sus-TBC1D15 belong to the same subfamily of TBC domain-containing proteins, and their GAP-domain structures are highly similar. This demonstrates the evolutionary conservation of the TBC1D15 protein family. Meanwhile, the newly determined crystal structures display new variations compared to the structures of yeast Gyp1p Rab GAP domain and TBC1D1. GAP assays show that Shark and Sus GAPs both have higher catalytic activity on Rab11a·GTP than Rab7a·GTP, which differs from the previous study. We also demonstrated the importance of arginine and glutamine on the catalytic sites of Shark GAP and Sus GAP. When arginine and glutamine are changed to alanine or lysine, the activities of Shark GAP and Sus GAP are lost.
- College of Life Science, Zhejiang Sci-Tech University, Hangzhou, 310018, China.
Organizational Affiliation: