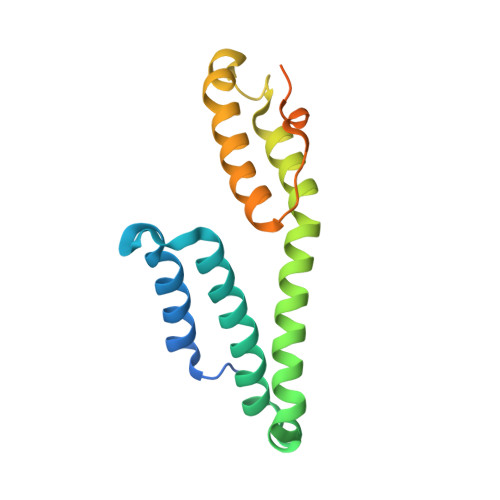
Malaria parasite CelTOS targets the inner leaflet of cell membranes for pore-dependent disruption.
Jimah, J.R., Salinas, N.D., Sala-Rabanal, M., Jones, N.G., Sibley, L.D., Nichols, C.G., Schlesinger, P.H., Tolia, N.H.(2016) Elife 5
- PubMed: 27906127 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.20621
- Primary Citation Related Structures:
5TSZ - PubMed Abstract:
Apicomplexan parasites contain a conserved protein CelTOS that, in malaria parasites, is essential for traversal of cells within the mammalian host and arthropod vector. However, the molecular role of CelTOS is unknown because it lacks sequence similarity to proteins of known function. Here, we determined the crystal structure of CelTOS and discovered CelTOS resembles proteins that bind to and disrupt membranes. In contrast to known membrane disruptors, CelTOS has a distinct architecture, specifically binds phosphatidic acid commonly present within the inner leaflet of plasma membranes, and potently disrupts liposomes composed of phosphatidic acid by forming pores. Microinjection of CelTOS into cells resulted in observable membrane damage. Therefore, CelTOS is unique as it achieves nearly universal inner leaflet cellular activity to enable the exit of parasites from cells during traversal. By providing novel molecular insight into cell traversal by apicomplexan parasites, our work facilitates the design of therapeutics against global pathogens.
- Department of Molecular Microbiology, Washington University School of Medicine, Saint Louis, United States.
Organizational Affiliation: