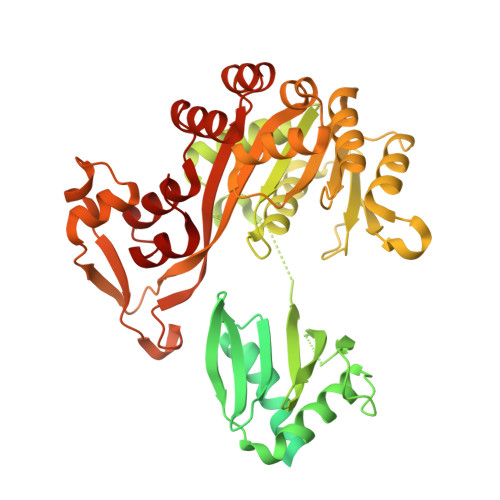
The molecular mechanism of the type IVa pilus motors.
McCallum, M., Tammam, S., Khan, A., Burrows, L.L., Howell, P.L.(2017) Nat Commun 8: 15091-15091
- PubMed: 28474682 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms15091
- Primary Citation Related Structures:
5TSG, 5TSH - PubMed Abstract:
Type IVa pili are protein filaments essential for virulence in many bacterial pathogens; they extend and retract from the surface of bacterial cells to pull the bacteria forward. The motor ATPase PilB powers pilus assembly. Here we report the structures of the core ATPase domains of Geobacter metallireducens PilB bound to ADP and the non-hydrolysable ATP analogue, AMP-PNP, at 3.4 and 2.3 Å resolution, respectively. These structures reveal important differences in nucleotide binding between chains. Analysis of these differences reveals the sequential turnover of nucleotide, and the corresponding domain movements. Our data suggest a clockwise rotation of the central sub-pores of PilB, which through interactions with PilC, would support the assembly of a right-handed helical pilus. Our analysis also suggests a counterclockwise rotation of the C2 symmetric PilT that would enable right-handed pilus disassembly. The proposed model provides insight into how this family of ATPases can power pilus extension and retraction.
- Department of Biochemistry, University of Toronto, Toronto, Ontario, Canada M5S 1A8.
Organizational Affiliation: