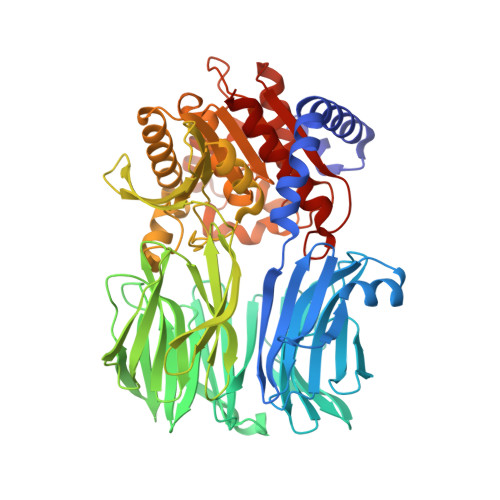
Crystal Structure and Conformational Dynamics of Pyrococcus furiosus Prolyl Oligopeptidase.
Ellis-Guardiola, K., Rui, H., Beckner, R.L., Srivastava, P., Sukumar, N., Roux, B., Lewis, J.C.(2019) Biochemistry 58: 1616-1626
- PubMed: 30786206
- DOI: https://doi.org/10.1021/acs.biochem.9b00031
- Primary Citation of Related Structures:
5T88, 6CAN - PubMed Abstract:
Enzymes in the prolyl oligopeptidase family possess unique structures and substrate specificities that are important for their biological activity and for potential biocatalytic applications. The crystal structures of Pyrococcus furiosus ( Pfu) prolyl oligopeptidase (POP) and the corresponding S477C mutant were determined to 1.9 and 2.2 Å resolution, respectively. The wild type enzyme crystallized in an open conformation, indicating that this state is readily accessible, and it contained bound chloride ions and a prolylproline ligand. These structures were used as starting points for molecular dynamics simulations of Pfu POP conformational dynamics. The simulations showed that large-scale domain opening and closing occurred spontaneously, providing facile substrate access to the active site. Movement of the loop containing the catalytically essential histidine into a conformation similar to those found in structures with fully formed catalytic triads also occurred. This movement was modulated by chloride binding, providing a rationale for experimentally observed activation of POP peptidase catalysis by chloride. Thus, the structures and simulations reported in this study, combined with existing biochemical data, provide a number of insights into POP catalysis.
- Department of Chemistry , University of Chicago , Chicago , Illinois 60637 , United States.
Organizational Affiliation: