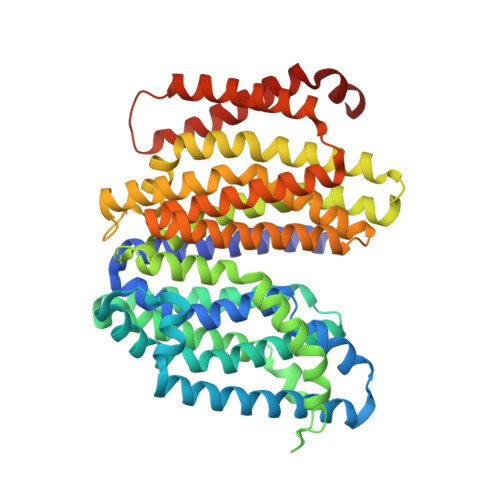
Crystal structure of the MOP flippase MurJ in an inward-facing conformation.
Kuk, A.C., Mashalidis, E.H., Lee, S.Y.(2017) Nat Struct Mol Biol 24: 171-176
- PubMed: 28024149 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.3346
- Primary Citation Related Structures:
5T77 - PubMed Abstract:
Peptidoglycan (PG) protects bacteria from osmotic lysis, and its biogenesis is a key antibiotic target. A central step in PG biosynthesis is flipping of the lipid-linked PG precursor lipid II across the cytoplasmic membrane for subsequent incorporation into PG. MurJ, part of the multidrug/oligosaccharidyl-lipid/polysaccharide (MOP) transporter superfamily, was recently shown to carry out this process. However, understanding of how MurJ flips lipid II, and of how MOP transporters operate in general, remains limited due to a lack of structural information. Here we present a crystal structure of MurJ from Thermosipho africanus in an inward-facing conformation at 2.0-Å resolution. A hydrophobic groove is formed by two C-terminal transmembrane helices, which leads into a large central cavity that is mostly cationic. Our studies not only provide the first structural glimpse of MurJ but also suggest that alternating access is important for MurJ function, which may be applicable to other MOP superfamily transporters.
- Department of Biochemistry, Duke University School of Medicine, Durham, North Carolina, USA.
Organizational Affiliation: