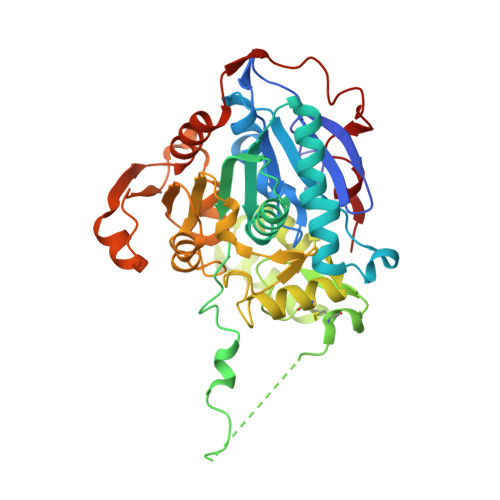
Structure of the Catalytic Domain of the Class I Polyhydroxybutyrate Synthase from Cupriavidus necator.
Wittenborn, E.C., Jost, M., Wei, Y., Stubbe, J., Drennan, C.L.(2016) J Biological Chem 291: 25264-25277
- PubMed: 27742839 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M116.756833
- Primary Citation Related Structures:
5T6O - PubMed Abstract:
Polyhydroxybutyrate synthase (PhaC) catalyzes the polymerization of 3-(R)-hydroxybutyryl-coenzyme A as a means of carbon storage in many bacteria. The resulting polymers can be used to make biodegradable materials with properties similar to those of thermoplastics and are an environmentally friendly alternative to traditional petroleum-based plastics. A full biochemical and mechanistic understanding of this process has been hindered in part by a lack of structural information on PhaC. Here we present the first structure of the catalytic domain (residues 201-589) of the class I PhaC from Cupriavidus necator (formerly Ralstonia eutropha) to 1.80 Å resolution. We observe a symmetrical dimeric architecture in which the active site of each monomer is separated from the other by ∼33 Å across an extensive dimer interface, suggesting a mechanism in which polyhydroxybutyrate biosynthesis occurs at a single active site. The structure additionally highlights key side chain interactions within the active site that play likely roles in facilitating catalysis, leading to the proposal of a modified mechanistic scheme involving two distinct roles for the active site histidine. We also identify putative substrate entrance and product egress routes within the enzyme, which are discussed in the context of previously reported biochemical observations. Our structure lays a foundation for further biochemical and structural characterization of PhaC, which could assist in engineering efforts for the production of eco-friendly materials.
- From the Departments of Chemistry.
Organizational Affiliation: