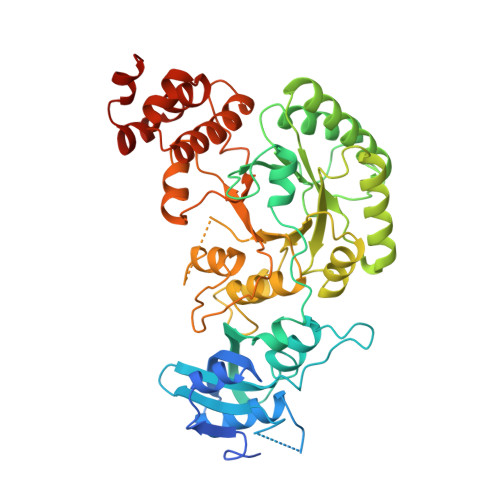
Crystallographic refinement and structure of ribulose-1,5-bisphosphate carboxylase from Rhodospirillum rubrum at 1.7 A resolution.
Schneider, G., Lindqvist, Y., Lundqvist, T.(1990) J Mol Biology 211: 989-1008
- PubMed: 2107319 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(90)90088-4
- Primary Citation Related Structures:
5RUB - PubMed Abstract:
The amino acid sequence of ribulose-1,5-bisphosphate carboxylase/oxygenase from Rhodospirillum rubrum has been fitted to the electron density maps. The resulting protein model has been refined to a nominal resolution of 1.7 A using the constrained-restrained least-squares refinement program of Sussman and the restrained least-squares refinement program of Hendrickson & Konnert. The crystallographic refinement, based on 76,452 reflections with F greater than sigma (F) in the resolution range 5.5 to 1.7 A resulted in a crystallographic R-factor of 18.0%. The asymmetric unit contains one dimeric ribulose-1,5-biphosphate carboxylase molecule, consisting of 869 amino acid residues and 736 water molecules. The geometry of the refined model is close to ideal, with root-mean-square deviations of 0.018 A in bond lengths and 2.7 degrees in bond angles. Two loop regions, comprising residues 54 to 63 and 324 to 335, and the last ten amino acid residues at the C terminus are disordered in our crystals. The expected trimodal distribution is obtained for the side-chain chi 1-angles with a marked preference for staggered conformation. The hydrogen-bonding pattern in the N-terminal beta-sheet and the parallel sheet in the beta/alpha-barrel is described. A number of hydrogen bonds and salt bridges are involved in domain-domain and subunit-subunit interactions. The subunit-subunit interface in the dimer covers an area of 2800 A2. Considerable deviations from the local 2-fold symmetry are found at both the N terminus (residues 2 to 5) and the C terminus (residues 422 to 457). Furthermore, loop 8 in the beta/alpha-barrel domain has a different conformation in the two subunits. A number of amino acid side-chains have different conformations in the two subunits. Most of these residues are located at the surface of the protein. An analysis of the individual temperature factors indicates a high mobility of the C-terminal region and for some of the loops at the active site. The positions and B-factors for 736 solvent sites have been refined (average B: 45.9 A2). Most of the solvent molecules are bound as clusters to the protein. The active site of the enzyme, especially the environment of the activator Lys191 in the non-activated enzyme is described. Crystallographic refinement at 1.7 A resolution clearly revealed the presence of a cis-proline at the active site. This residue is part of the highly conserved region Lys166-Pro167-Lys168.
- Swedish University of Agricultural Sciences, Uppsala Biomedical Centre, Department of Molecular Biology.
Organizational Affiliation: