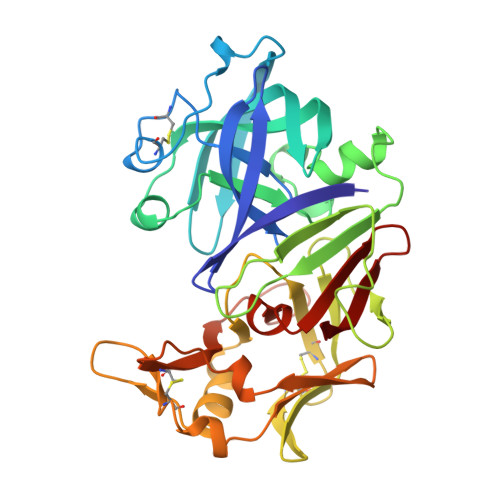
X-ray analyses of aspartic proteinases. II. Three-dimensional structure of the hexagonal crystal form of porcine pepsin at 2.3 A resolution.
Cooper, J.B., Khan, G., Taylor, G., Tickle, I.J., Blundell, T.L.(1990) J Mol Biology 214: 199-222
- PubMed: 2115088 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(90)90156-G
- Primary Citation Related Structures:
5PEP - PubMed Abstract:
The molecular structure of the hexagonal crystal form of porcine pepsin (EC 3.4.23.1), an aspartic proteinase from the gastric mucosa, has been determined by molecular replacement using the fungal enzyme, penicillopepsin (EC 3.4.23.6), as the search model. This defined the space group as P6522 and refinement led to an R-factor of 0.190 at 2.3 A resolution. The positions of 2425 non-hydrogen protein atoms in 326 residues have been determined and the model contains 371 water molecules. The structure is bilobal, consisting of two predominantly beta-sheet lobes which, as in other aspartic proteinases, are related by a pseudo 2-fold axis. The strands of the mixed beta-sheets (1N and 1C) of each lobe are related by an intra-lobe topological 2-fold symmetry. Two further beta-sheets, 2N and 2C, are each composed of two topologically related beta-hairpins folded below the 1N and 1C sheets. A further six-stranded sheet (3) spans the two lobes and forms a structure resembling an arch upon which the four other sheets reside. The interface between sheets 1N and 1C forms the catalytic centre consisting of absolutely conserved aspartate residues 32 and 215, which are shielded from solvent by a beta-hairpin loop (75 to 78). The crystal structure of a mammalian aspartic proteinase indicates that interactions with substrate may be more extensive on the prime side of the active site cleft than in the fungal enzymes and involve Tyr189 and the loop 290 to 295, perhaps contributing to the transpeptidase activity of pepsin and the specificity of the renins. Comparison with the high-resolution structure of pepsinogen gives a root-mean-square deviation of 0.9 A and reveals that, in addition to local rearrangement at the active site, there appears to be a rigid group movement of part of the C-terminal lobe of pepsin towards the cleft on activation. A large proportion of the absolutely conserved residues in aspartic proteinases are polar and buried. An examination of the pepsin structure reveals that these side-chains are involved in hydrogen-bond interactions with either the main chain of the protein or other conserved side-chains of the enzyme or propart.
- Department of Crystallography, Birkbeck College, London, U.K.
Organizational Affiliation: