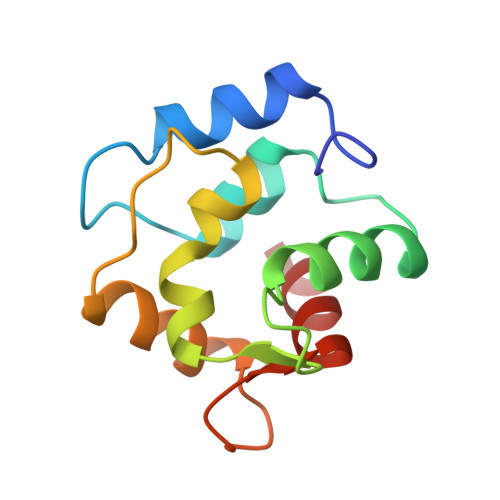
Crystal structure of the unique parvalbumin component from muscle of the leopard shark (Triakis semifasciata). The first X-ray study of an alpha-parvalbumin.
Roquet, F., Declercq, J.P., Tinant, B., Rambaud, J., Parello, J.(1992) J Mol Biology 223: 705-720
- PubMed: 1542115
- DOI: https://doi.org/10.1016/0022-2836(92)90985-s
- Primary Citation Related Structures:
5PAL - PubMed Abstract:
The three-dimensional structure of parvalbumin from leopard shark (Triakis semifasciata) with 109 amino acid residues (alpha-series) is described at 1.54 A resolution. Crystals were grown at 20 degrees C from 2.9 M-potassium/sodium phosphate solutions at pH 5.6. The space group is P3(1)21 and unit cell dimensions are a = b = 32.12 A and c = 149.0 A. The structure has been solved by the molecular replacement method using pike 4.10 parvalbumin as a model. The final structure refinement resulted in an R-factor of 17.3% for 11,363 independent reflections at 1.54 A resolution. The shark parvalbumin shows the main features of all parvalbumins: the folding of the chain including six alpha-helices, the salt bridge between Arg75 and Glu81, and the hydrophobic core. Compared to the structure of beta-parvalbumins from pike and carp, one main difference is observed: the chain is one residue longer and this additional residue, which extends the F helix, is involved through its C-terminal carboxylate group in a network of electrostatic contacts with two basic residues, His31 in the B helix and Lys36 in the BC segment. Furthermore, hydrogen bonds exist between the side-chains of Gln108 (F helix) and Tyr26 (B helix). There is therefore a "locking" of the tertiary structure through contacts between two sequentially distant regions in the protein and this is likely to contribute to making the stability of an alpha-parvalbumin higher in comparison to that of a beta-parvalbumin. The lengthening of the C-terminal F helix by one residue appears to be a major feature of alpha-parvalbumins in general, owing to the homologies of the amino acid sequences. Besides the lengthening of the C-terminal helix, the classification of the leopard shark parvalbumin in the alpha-series rests upon the observation of Lys13, Leu32, Glu61 and Val66. As this is the first crystal structure description of a parvalbumin from the alpha-phylogenetic lineage, it was hoped that it would clearly determine the presence or absence of a third cation binding site in parvalbumins belonging to the alpha-lineage. In beta-pike pI 4.10 parvalbumin, Asp61 participates as a direct ligand of a third site, the satellite of the CD site. In shark parvalbumin, as in nearly all alpha-parvalbumins, one finds Glu at position 61. Unfortunately, the conformation of the polar head of Glu61 cannot be inferred from the X-ray data.(ABSTRACT TRUNCATED AT 400 WORDS)
- Laboratoire de Chimie-Physique et de Cristallographie, Université Catholique de Louvain, Belgium.
Organizational Affiliation: