Crystal Structure of the Protein-Kinase A catalytic subunit from Criteculus Griseus in complex with compounds RKp013 and RKp190
Mueller, J.M., Kirschner, R., Heine, A., Geyer, A., Klebe, G.To be published.
Experimental Data Snapshot
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| cAMP-dependent protein kinase catalytic subunit alpha | 353 | Cricetulus griseus | Mutation(s): 0 Gene Names: PRKACA EC: 2.7.11.11 | 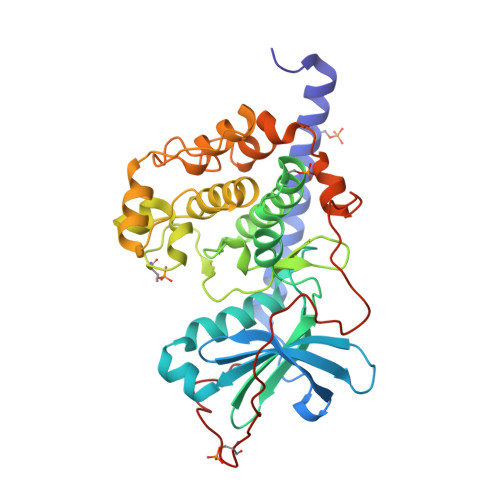 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P25321 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| cAMP-dependent protein kinase inhibitor | B [auth D] | 18 | Cricetulus griseus | Mutation(s): 0 | 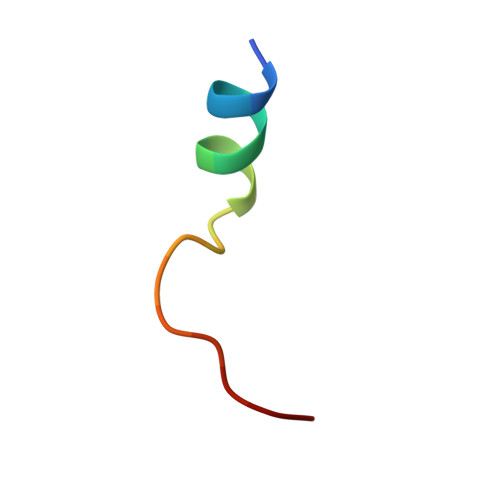 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | G3HK48 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| AO8 Download:Ideal Coordinates CCD File | C [auth A] | [2-[[2-(isoquinolin-5-ylsulfonylamino)ethylamino]methyl]phenyl]boronic acid C18 H20 B N3 O4 S KGPSPJNLVIXNTL-UHFFFAOYSA-N |  | ||
| Modified Residues 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| SEP Query on SEP | A | L-PEPTIDE LINKING | C3 H8 N O6 P |  | SER |
| TPO Query on TPO | A | L-PEPTIDE LINKING | C4 H10 N O6 P |  | THR |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 109.381 | α = 90 |
| b = 59.167 | β = 90 |
| c = 74.223 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| XDS | data scaling |
| PHASER | phasing |
| Coot | model building |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Loewe-Corporation | Germany | -- |