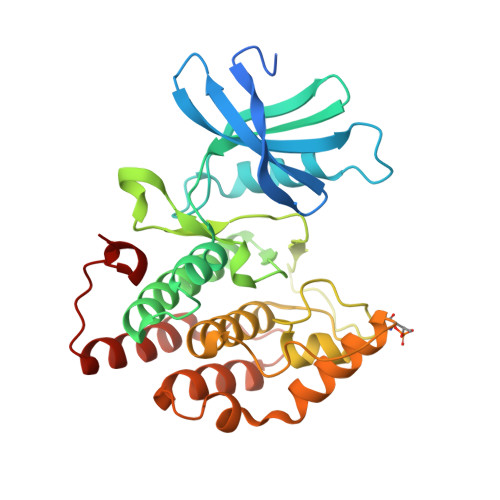
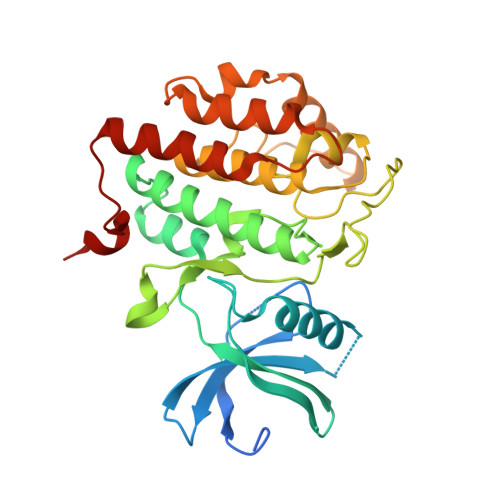
Discovery of Inhibitor of Wnt Production 2 (IWP-2) and Related Compounds As Selective ATP-Competitive Inhibitors of Casein Kinase 1 (CK1) delta / epsilon.
Garcia-Reyes, B., Witt, L., Jansen, B., Karasu, E., Gehring, T., Leban, J., Henne-Bruns, D., Pichlo, C., Brunstein, E., Baumann, U., Wesseler, F., Rathmer, B., Schade, D., Peifer, C., Knippschild, U.(2018) J Med Chem 61: 4087-4102
- PubMed: 29630366 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.8b00095
- Primary Citation Related Structures:
5OKT - PubMed Abstract:
Inhibitors of Wnt production (IWPs) are known antagonists of the Wnt pathway, targeting the membrane-bound O-acyltransferase porcupine (Porcn) and thus preventing a crucial Wnt ligand palmitoylation. Since IWPs show structural similarities to benzimidazole-based CK1 inhibitors, we hypothesized that IWPs could also inhibit CK1 isoforms. Molecular modeling revealed a plausible binding mode of IWP-2 in the ATP binding pocket of CK1δ which was confirmed by X-ray analysis. In vitro kinase assays demonstrated IWPs to be ATP-competitive inhibitors of wt CK1δ. IWPs also strongly inhibited the gatekeeper mutant M82F CK1δ. When profiled in a panel of 320 kinases, IWP-2 specifically inhibited CK1δ. IWP-2 and IWP-4 also inhibited the viability of various cancer cell lines. By a medicinal chemistry approach, we developed improved IWP-derived CK1 inhibitors. Our results suggest that the effects of IWPs are not limited to Porcn, but also might influence CK1δ/ε-related pathways.
- Department of General and Visceral Surgery , Ulm University Hospital , Albert-Einstein-Allee 23 , D-89081 Ulm , Germany.
Organizational Affiliation: