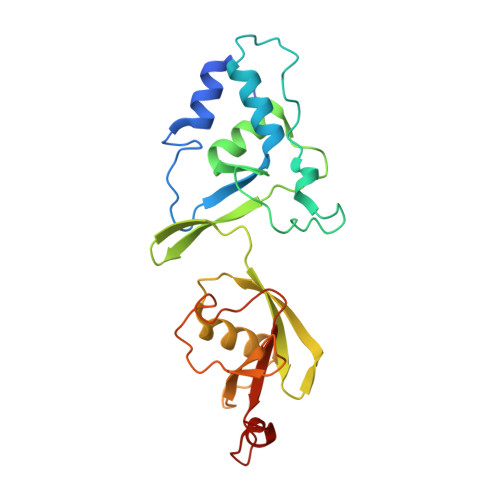
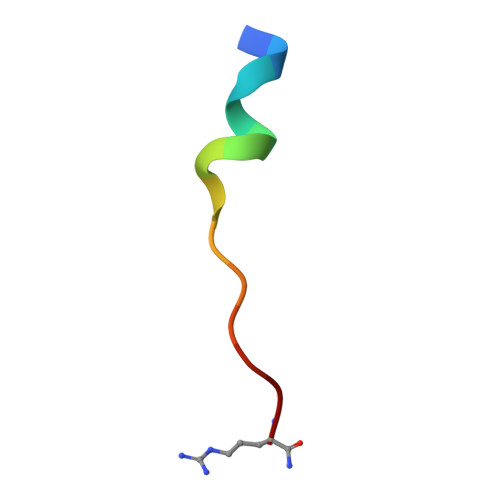
Discovery of peptide ligands targeting a specific ubiquitin-like domain-binding site in the deubiquitinase USP11.
Spiliotopoulos, A., Blokpoel Ferreras, L., Densham, R.M., Caulton, S.G., Maddison, B.C., Morris, J.R., Dixon, J.E., Gough, K.C., Dreveny, I.(2019) J Biological Chem 294: 424-436
- PubMed: 30373771 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA118.004469
- Primary Citation Related Structures:
5OK6 - PubMed Abstract:
Ubiquitin-specific proteases (USPs) reverse ubiquitination and regulate virtually all cellular processes. Defined noncatalytic domains in USP4 and USP15 are known to interact with E3 ligases and substrate recruitment factors. No such interactions have been reported for these domains in the paralog USP11, a key regulator of DNA double-strand break repair by homologous recombination. We hypothesized that USP11 domains adjacent to its protease domain harbor unique peptide-binding sites. Here, using a next-generation phage display (NGPD) strategy, combining phage display library screening with next-generation sequencing, we discovered unique USP11-interacting peptide motifs. Isothermal titration calorimetry disclosed that the highest affinity peptides ( K D of ∼10 μm) exhibit exclusive selectivity for USP11 over USP4 and USP15 in vitro Furthermore, a crystal structure of a USP11-peptide complex revealed a previously unknown binding site in USP11's noncatalytic ubiquitin-like (UBL) region. This site interacted with a helical motif and is absent in USP4 and USP15. Reporter assays using USP11-WT versus a binding pocket-deficient double mutant disclosed that this binding site modulates USP11's function in homologous recombination-mediated DNA repair. The highest affinity USP11 peptide binder fused to a cellular delivery sequence induced significant nuclear localization and cell cycle arrest in S phase, affecting the viability of different mammalian cell lines. The USP11 peptide ligands and the paralog-specific functional site in USP11 identified here provide a framework for the development of new biochemical tools and therapeutic agents. We propose that an NGPD-based strategy for identifying interacting peptides may be applied also to other cellular targets.
- From the Centre for Biomolecular Sciences, School of Pharmacy, University of Nottingham, Nottingham NG7 2RD.
Organizational Affiliation: