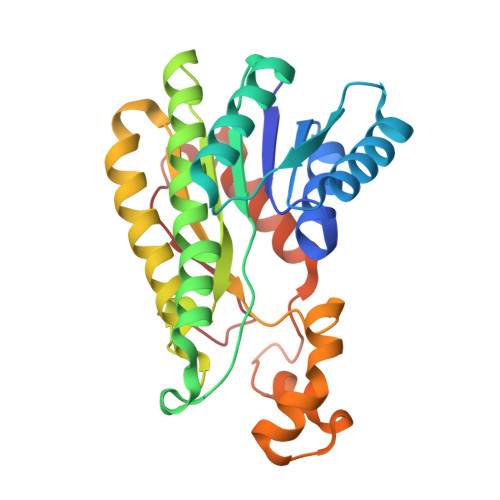
Structural and functional analysis of Erwinia amylovora SrlD. The first crystal structure of a sorbitol-6-phosphate 2-dehydrogenase.
Salomone-Stagni, M., Bartho, J.D., Kalita, E., Rejzek, M., Field, R.A., Bellini, D., Walsh, M.A., Benini, S.(2018) J Struct Biol 203: 109-119
- PubMed: 29605571
- DOI: https://doi.org/10.1016/j.jsb.2018.03.010
- Primary Citation of Related Structures:
5O3Z - PubMed Abstract:
Sorbitol-6-phosphate 2-dehydrogenases (S6PDH) catalyze the interconversion of d-sorbitol 6-phosphate to d-fructose 6-phosphate. In the plant pathogen Erwinia amylovora the S6PDH SrlD is used by the bacterium to utilize sorbitol, which is used for carbohydrate transport in the host plants belonging to the Amygdaloideae subfamily (e.g., apple, pear, and quince). We have determined the crystal structure of S6PDH SrlD at 1.84 Å resolution, which is the first structure of an EC 1.1.1.140 enzyme. Kinetic data show that SrlD is much faster at oxidizing d-sorbitol 6-phosphate than in reducing d-fructose 6-phosphate, however, equilibrium analysis revealed that only part of the d-sorbitol 6-phosphate present in the in vitro environment is converted into d-fructose 6-phosphate. The comparison of the structures of SrlD and Rhodobacter sphaeroides sorbitol dehydrogenase showed that the tetrameric quaternary structure, the catalytic residues and a conserved aspartate residue that confers specificity for NAD + over NADP + are preserved. Analysis of the SrlD cofactor and substrate binding sites identified residues important for the formation of the complex with cofactor and substrate and in particular the role of Lys42 in selectivity towards the phospho-substrate. The comparison of SrlD backbone with the backbone of 302 short-chain dehydrogenases/reductases showed the conservation of the protein core and identified the variable parts. The SrlD sequence was compared with 500 S6PDH sequences selected by homology revealing that the C-terminal part is more conserved than the N-terminal, the consensus of the catalytic tetrad (Y[SN]AGXA) and a not previously described consensus for the NAD(H) binding.
- Bioorganic Chemistry and Bio-Crystallography Laboratory (B(2)Cl), Faculty of Science and Technology, Free University of Bolzano, Piazza Università 5, 39100 Bolzano, Italy.
Organizational Affiliation: