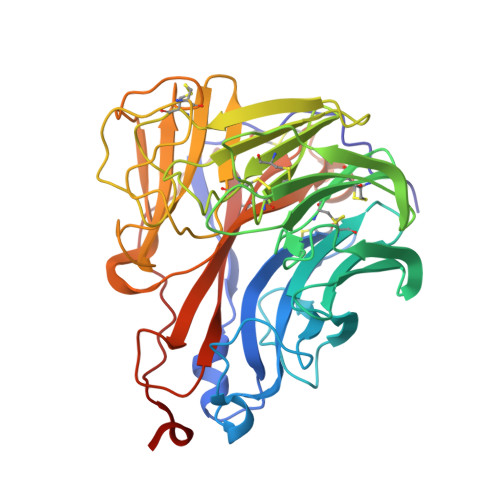
Kinetic, Thermodynamic, and Structural Analysis of Drug Resistance Mutations in Neuraminidase from the 2009 Pandemic Influenza Virus.
Pokorna, J., Pachl, P., Karlukova, E., Hejdanek, J., Rezacova, P., Machara, A., Hudlicky, J., Konvalinka, J., Kozisek, M.(2018) Viruses 10
- PubMed: 29933553
- DOI: https://doi.org/10.3390/v10070339
- Primary Citation of Related Structures:
5NWE, 5NZ4, 5NZE, 5NZF, 5NZN - PubMed Abstract:
Neuraminidase is the main target for current influenza drugs. Reduced susceptibility to oseltamivir, the most widely prescribed neuraminidase inhibitor, has been repeatedly reported. The resistance substitutions I223V and S247N, alone or in combination with the major oseltamivir-resistance mutation H275Y, have been observed in 2009 pandemic H1N1 viruses. We overexpressed and purified the ectodomain of wild-type neuraminidase from the A/California/07/2009 (H1N1) influenza virus, as well as variants containing H275Y, I223V, and S247N single mutations and H275Y/I223V and H275Y/S247N double mutations. We performed enzymological and thermodynamic analyses and structurally examined the resistance mechanism. Our results reveal that the I223V or S247N substitution alone confers only a moderate reduction in oseltamivir affinity. In contrast, the major oseltamivir resistance mutation H275Y causes a significant decrease in the enzyme’s ability to bind this drug. Combination of H275Y with an I223V or S247N mutation results in extreme impairment of oseltamivir’s inhibition potency. Our structural analyses revealed that the H275Y substitution has a major effect on the oseltamivir binding pose within the active site while the influence of other studied mutations is much less prominent. Our crystal structures also helped explain the augmenting effect on resistance of combining H275Y with both substitutions.
- The Czech Academy of Sciences, Institute of Organic Chemistry and Biochemistry, Flemingovo n. 2, 166 10 Prague 6, Czech Republic. jana.pokorna@uochb.cas.cz.
Organizational Affiliation: