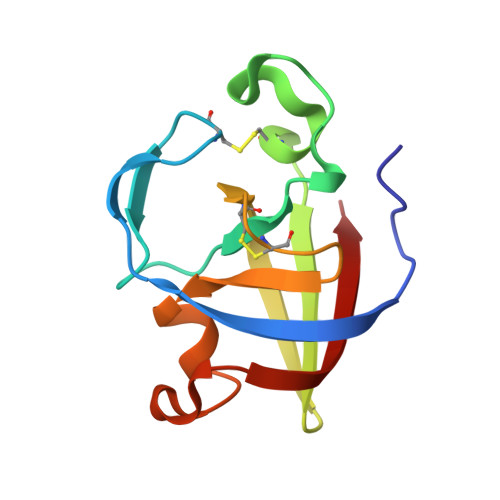
Illuminating structure and acyl donor sites of a physiological transglutaminase substrate from Streptomyces mobaraensis.
Juettner, N.E., Schmelz, S., Bogen, J.P., Happel, D., Fessner, W.D., Pfeifer, F., Fuchsbauer, H.L., Scrima, A.(2018) Protein Sci 27: 910-922
- PubMed: 29430769 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3388
- Primary Citation Related Structures:
5NTB - PubMed Abstract:
Transglutaminase from Streptomyces mobaraensis (MTG) has become a powerful tool to covalently and highly specifically link functional amines to glutamine donor sites of therapeutic proteins. However, details regarding the mechanism of substrate recognition and interaction of the enzyme with proteinaceous substrates still remain mostly elusive. We have determined the crystal structure of the Streptomyces papain inhibitory protein (SPI p ), a substrate of MTG, to study the influence of various substrate amino acids on positioning glutamine to the active site of MTG. SPI p exhibits a rigid, thermo-resistant double-psi-beta-barrel fold that is stabilized by two cysteine bridges. Incorporation of biotin cadaverine identified Gln-6 as the only amine acceptor site on SPI p accessible for MTG. Substitution of Lys-7 demonstrated that small and hydrophobic residues in close proximity to Gln-6 favor MTG-mediated modification and are likely to facilitate introduction of the substrate into the front vestibule of MTG. Moreover, exchange of various surface residues of SPI p for arginine and glutamate/aspartate outside the glutamine donor region influences the efficiency of modification by MTG. These results suggest the occurrence of charged contact areas between MTG and the acyl donor substrates beyond the front vestibule, and pave the way for protein engineering approaches to improve the properties of artificial MTG-substrates used in biomedical applications.
- The Department of Chemical Engineering and Biotechnology, University of Applied Sciences of Darmstadt, Darmstadt, Germany.
Organizational Affiliation: