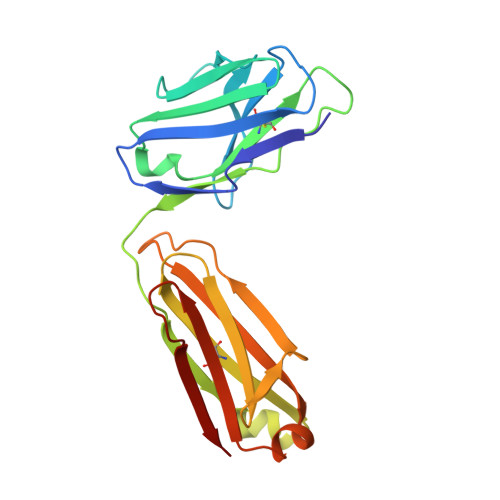
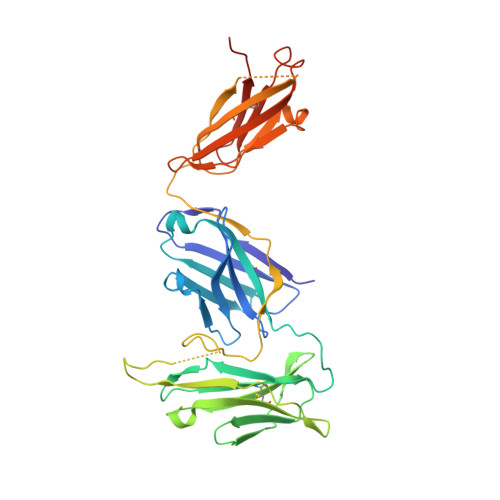
The structure of a LAIR1-containing human antibody reveals a novel mechanism of antigen recognition.
Hsieh, F.L., Higgins, M.K.(2017) Elife 6
- PubMed: 28527239 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.27311
- Primary Citation Related Structures:
5NST - PubMed Abstract:
Antibodies are critical components of the human adaptive immune system, providing versatile scaffolds to display diverse antigen-binding surfaces. Nevertheless, most antibodies have similar architectures, with the variable immunoglobulin domains of the heavy and light chain each providing three hypervariable loops, which are varied to generate diversity. The recent identification of a novel class of antibody in humans from malaria endemic regions of Africa was therefore surprising as one hypervariable loop contains the entire collagen-binding domain of human LAIR1. Here, we present the structure of the Fab fragment of such an antibody. We show that its antigen-binding site has adopted an architecture that positions LAIR1, while itself being occluded. This therefore represents a novel means of antigen recognition, in which the Fab fragment of an antibody acts as an adaptor, linking a human protein insert with antigen-binding potential to the constant antibody regions which mediate immune cell recruitment.
- Department of Biochemistry, University of Oxford, Oxford, United Kingdom.
Organizational Affiliation: