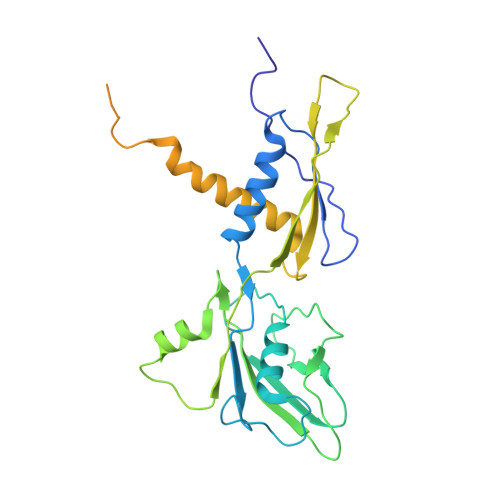
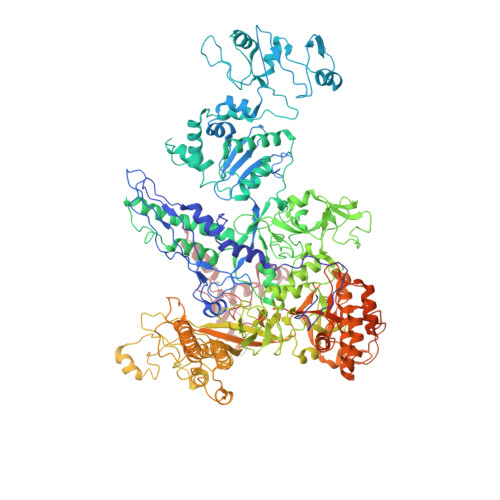
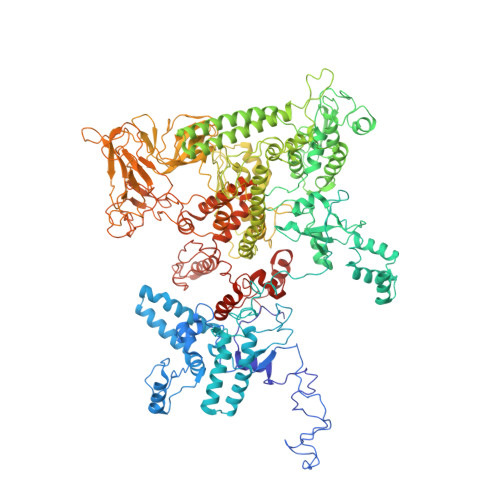
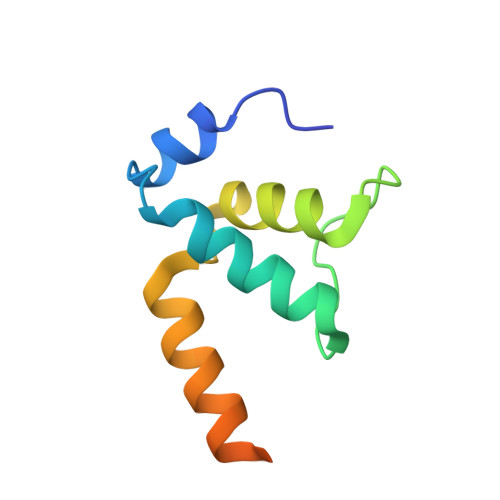
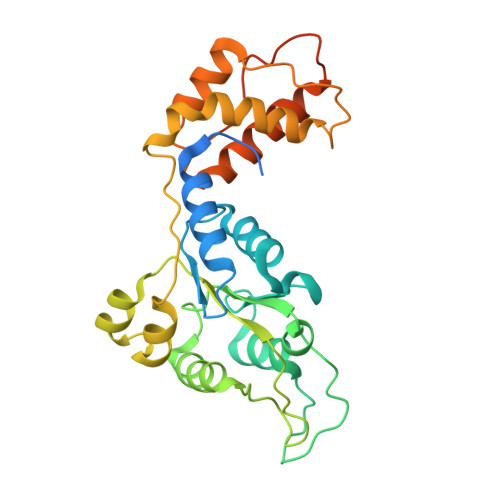
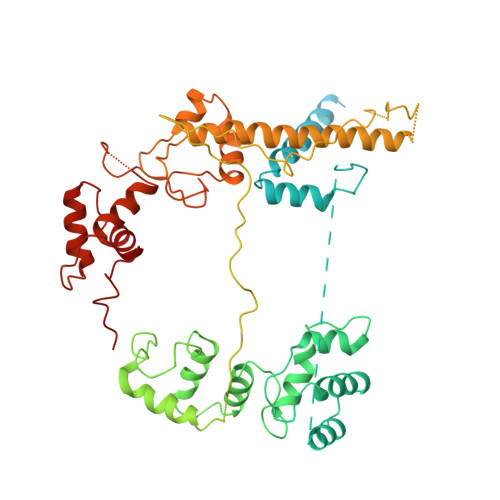
Structures of RNA Polymerase Closed and Intermediate Complexes Reveal Mechanisms of DNA Opening and Transcription Initiation.
Glyde, R., Ye, F., Darbari, V.C., Zhang, N., Buck, M., Zhang, X.(2017) Mol Cell 67: 106-116.e4
- PubMed: 28579332 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2017.05.010
- Primary Citation Related Structures:
5NSR, 5NSS - PubMed Abstract:
Gene transcription is carried out by RNA polymerases (RNAPs). For transcription to occur, the closed promoter complex (RPc), where DNA is double stranded, must isomerize into an open promoter complex (RPo), where the DNA is melted out into a transcription bubble and the single-stranded template DNA is delivered to the RNAP active site. Using a bacterial RNAP containing the alternative σ 54 factor and cryoelectron microscopy, we determined structures of RPc and the activator-bound intermediate complex en route to RPo at 3.8 and 5.8 Å. Our structures show how RNAP-σ 54 interacts with promoter DNA to initiate the DNA distortions required for transcription bubble formation, and how the activator interacts with RPc, leading to significant conformational changes in RNAP and σ 54 that promote RPo formation. We propose that DNA melting is an active process initiated in RPc and that the RNAP conformations of intermediates are significantly different from that of RPc and RPo.
- Section of Structural Biology, Department of Medicine, Imperial College London, London SW7 2AZ, UK.
Organizational Affiliation: