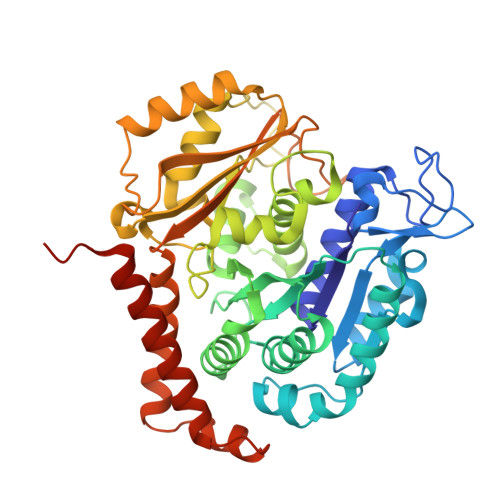
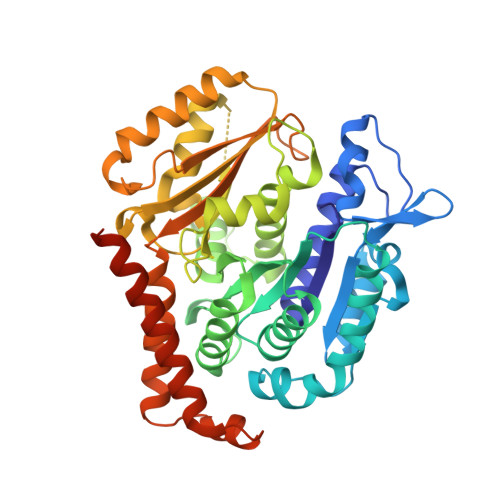
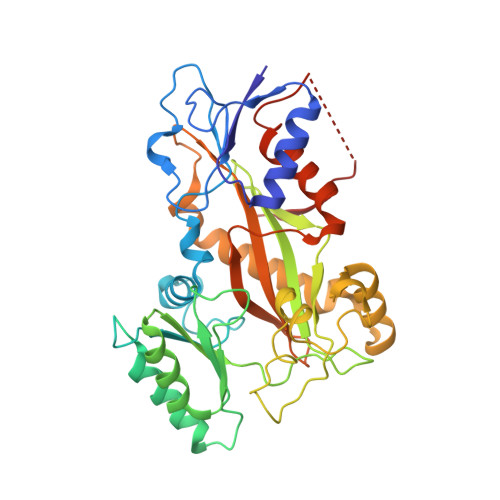
Zampanolide Binding to Tubulin Indicates Cross-Talk of Taxane Site with Colchicine and Nucleotide Sites.
Field, J.J., Pera, B., Gallego, J.E., Calvo, E., Rodriguez-Salarichs, J., Saez-Calvo, G., Zuwerra, D., Jordi, M., Andreu, J.M., Prota, A.E., Menchon, G., Miller, J.H., Altmann, K.H., Diaz, J.F.(2018) J Nat Prod 81: 494-505
- PubMed: 29023132 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jnatprod.7b00704
- Primary Citation Related Structures:
5NFZ, 5NG1 - PubMed Abstract:
The marine natural product zampanolide and analogues thereof constitute a new chemotype of taxoid site microtubule-stabilizing agents with a covalent mechanism of action. Zampanolide-ligated tubulin has the switch-activation loop (M-loop) in the assembly prone form and, thus, represents an assembly activated state of the protein. In this study, we have characterized the biochemical properties of the covalently modified, activated tubulin dimer, and we have determined the effect of zampanolide on tubulin association and the binding of tubulin ligands at other binding sites. Tubulin activation by zampanolide does not affect its longitudinal oligomerization but does alter its lateral association properties. The covalent binding of zampanolide to β-tubulin affects both the colchicine site, causing a change of the quantum yield of the bound ligand, and the exchangeable nucleotide binding site, reducing the affinity for the nucleotide. While these global effects do not change the binding affinity of 2-methoxy-5-(2,3,4-trimethoxyphenyl)-2,4,6-cycloheptatrien-1-one (MTC) (a reversible binder of the colchicine site), the binding affinity of a fluorescent analogue of GTP (Mant-GTP) at the nucleotide E-site is reduced from 12 ± 2 × 10 5 M -1 in the case of unmodified tubulin to 1.4 ± 0.3 × 10 5 M -1 in the case of the zampanolide tubulin adduct, indicating signal transmission between the taxane site and the colchicine and nucleotide sites of β-tubulin.
- Centre for Biodiscovery, School of Biological Sciences , Victoria University of Wellington , Wellington 6012 , New Zealand.
Organizational Affiliation: