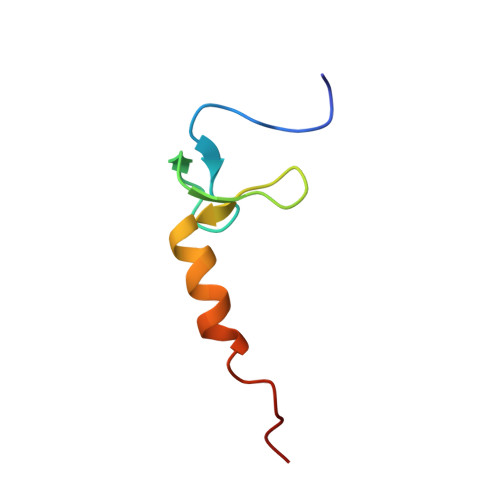
Structural and Functional Insights into Bacillus subtilis Sigma Factor Inhibitor, CsfB.
Martinez-Lumbreras, S., Alfano, C., Evans, N.J., Collins, K.M., Flanagan, K.A., Atkinson, R.A., Krysztofinska, E.M., Vydyanath, A., Jackter, J., Fixon-Owoo, S., Camp, A.H., Isaacson, R.L.(2018) Structure 26: 640-648.e5
- PubMed: 29526435 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2018.02.007
- Primary Citation Related Structures:
5N7Y - PubMed Abstract:
Global changes in bacterial gene expression can be orchestrated by the coordinated activation/deactivation of alternative sigma (σ) factor subunits of RNA polymerase. Sigma factors themselves are regulated in myriad ways, including via anti-sigma factors. Here, we have determined the solution structure of anti-sigma factor CsfB, responsible for inhibition of two alternative sigma factors, σ G and σ E , during spore formation by Bacillus subtilis. CsfB assembles into a symmetrical homodimer, with each monomer bound to a single Zn 2+ ion via a treble-clef zinc finger fold. Directed mutagenesis indicates that dimer formation is critical for CsfB-mediated inhibition of both σ G and σ E , and we have characterized these interactions in vitro. This work represents an advance in our understanding of how CsfB mediates inhibition of two alternative sigma factors to drive developmental gene expression in a bacterium.
- Department of Chemistry, King's College London, Britannia House, 7 Trinity Street, London SE1 1DB, UK.
Organizational Affiliation: