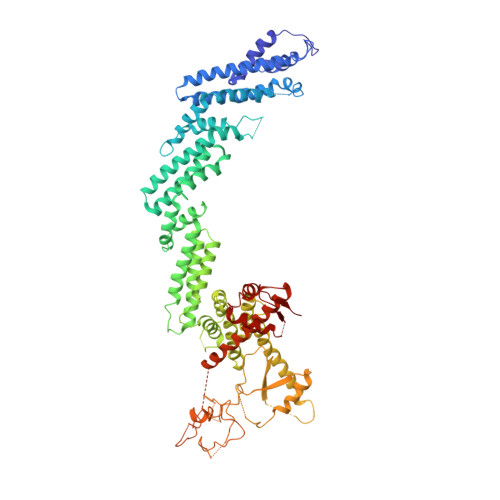
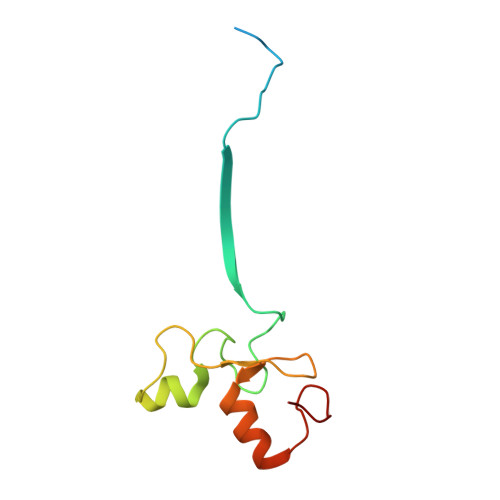
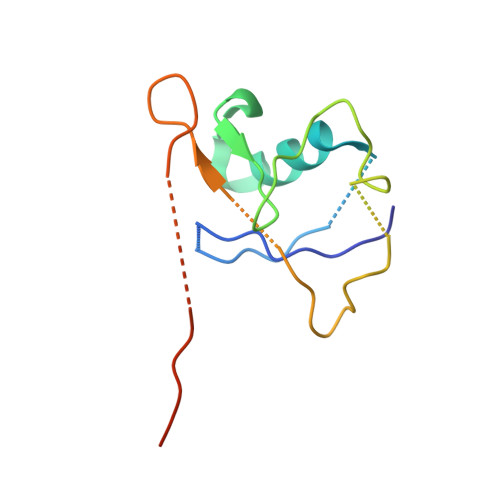
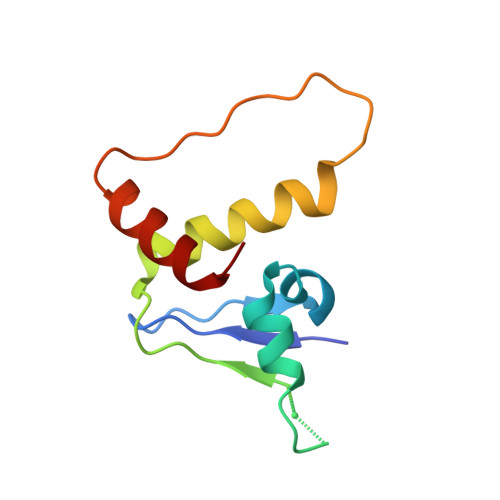
Crystal Structure of the Cul2-Rbx1-EloBC-VHL Ubiquitin Ligase Complex.
Cardote, T.A.F., Gadd, M.S., Ciulli, A.(2017) Structure 25: 901-911.e3
- PubMed: 28591624 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2017.04.009
- Primary Citation Related Structures:
5N4W - PubMed Abstract:
Cullin RING E3 ubiquitin ligases (CRLs) function in the ubiquitin proteasome system to catalyze the transfer of ubiquitin from E2 conjugating enzymes to specific substrate proteins. CRLs are large dynamic complexes and attractive drug targets for the development of small-molecule inhibitors and chemical inducers of protein degradation. The atomic details of whole CRL assembly and interactions that dictate subunit specificity remain elusive. Here we present the crystal structure of a pentameric CRL2 VHL complex, composed of Cul2, Rbx1, Elongin B, Elongin C, and pVHL. The structure traps a closed state of full-length Cul2 and a new pose of Rbx1 in a trajectory from closed to open conformation. We characterize hotspots and binding thermodynamics at the interface between Cul2 and pVHL-EloBC and identify mutations that contribute toward a selectivity switch for Cul2 versus Cul5 recognition. Our findings provide structural and biophysical insights into the whole Cul2 complex that could aid future drug targeting.
- Division of Biological Chemistry and Drug Discovery, School of Life Sciences, University of Dundee, Dow Street, Dundee DD1 5EH, UK.
Organizational Affiliation: