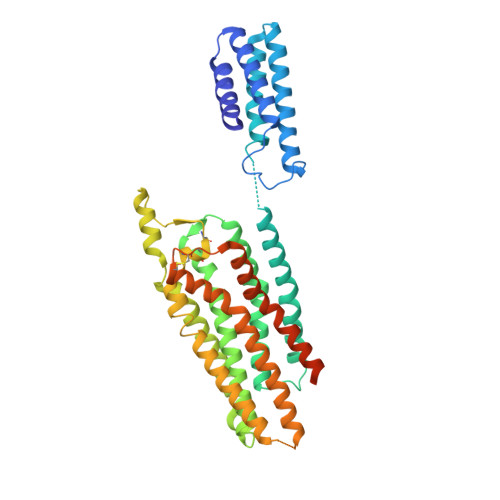
Structures of Human A1 and A2A Adenosine Receptors with Xanthines Reveal Determinants of Selectivity.
Cheng, R.K.Y., Segala, E., Robertson, N., Deflorian, F., Dore, A.S., Errey, J.C., Fiez-Vandal, C., Marshall, F.H., Cooke, R.M.(2017) Structure 25: 1275-1285.e4
- PubMed: 28712806
- DOI: https://doi.org/10.1016/j.str.2017.06.012
- Primary Citation of Related Structures:
5MZJ, 5MZP, 5N2R, 5N2S - PubMed Abstract:
The adenosine A 1 and A 2A receptors belong to the purinergic family of G protein-coupled receptors, and regulate diverse functions of the cardiovascular, respiratory, renal, inflammation, and CNS. Xanthines such as caffeine and theophylline are weak, non-selective antagonists of adenosine receptors. Here we report the structure of a thermostabilized human A 1 receptor at 3.3 Å resolution with PSB36, an A 1 -selective xanthine-based antagonist. This is compared with structures of the A 2A receptor with PSB36 (2.8 Å resolution), caffeine (2.1 Å), and theophylline (2.0 Å) to highlight features of ligand recognition which are common across xanthines. The structures of A 1 R and A 2A R were analyzed to identify the differences that are important selectivity determinants for xanthine ligands, and the role of T270 7.35 in A 1 R (M270 7.35 in A 2A R) in conferring selectivity was confirmed by mutagenesis. The structural differences confirmed to lead to selectivity can be utilized in the design of new subtype-selective A 1 R or A 2A R antagonists.
- Heptares Therapeutics Ltd, Biopark, Broadwater Road, Welwyn Garden City AL7 3AX, UK.
Organizational Affiliation: