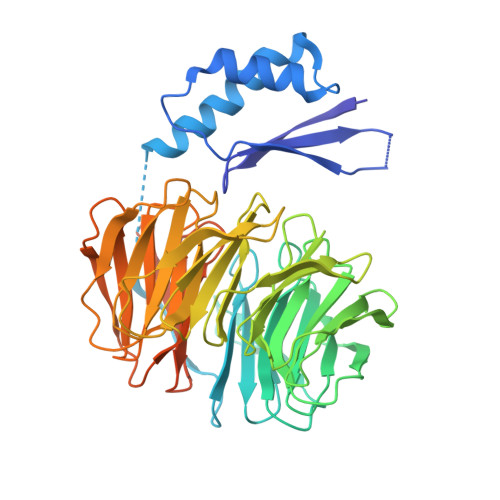
Structural basis of outer dynein arm intraflagellar transport by the transport adaptor protein ODA16 and the intraflagellar transport protein IFT46.
Taschner, M., Mourao, A., Awasthi, M., Basquin, J., Lorentzen, E.(2017) J Biological Chem 292: 7462-7473
- PubMed: 28298440 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M117.780155
- Primary Citation Related Structures:
5MZH - PubMed Abstract:
Motile cilia are found on unicellular organisms such as the green alga Chlamydomonas reinhardtii , on sperm cells, and on cells that line the trachea and fallopian tubes in mammals. The motility of cilia relies on a number of large protein complexes including the force-generating outer dynein arms (ODAs). The transport of ODAs into cilia has been previously shown to require the transport adaptor ODA16, as well as the intraflagellar transport (IFT) protein IFT46, but the molecular mechanism by which ODAs are recognized and transported into motile cilia is still unclear. Here, we determined the high-resolution crystal structure of C. reinhardtii ODA16 (CrODA16) and mapped the binding to IFT46 and ODAs. The CrODA16 structure revealed a small 80-residue N-terminal domain and a C-terminal 8-bladed β-propeller domain that are both required for the association with the N-terminal 147 residues of IFT46. The dissociation constant of the IFT46-ODA16 complex was 200 nm, demonstrating that CrODA16 associates with the IFT complex with an affinity comparable with that of the individual IFT subunits. Furthermore, we show, using ODAs extracted from the axonemes of C. reinhardtii , that the C-terminal β-propeller but not the N-terminal domain of CrODA16 is required for the interaction with ODAs. These data allowed us to present an architectural model for ODA16-mediated IFT of ODAs.
- From the Department of Molecular Biology and Genetics, Aarhus University, DK-8000 Aarhus C, Denmark.
Organizational Affiliation: