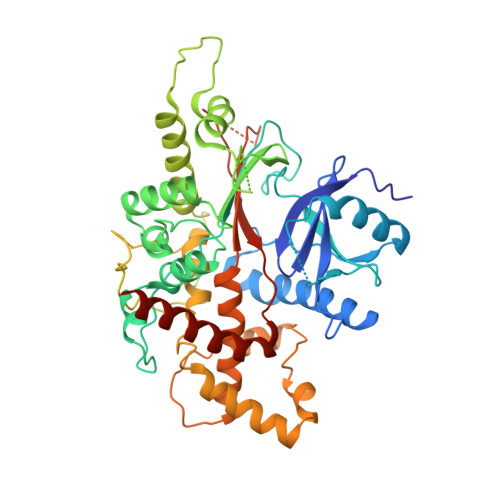
The crystal structure of mammalian inositol 1,3,4,5,6-pentakisphosphate 2-kinase reveals a new zinc-binding site and key features for protein function.
Franco-Echevarria, E., Sanz-Aparicio, J., Brearley, C.A., Gonzalez-Rubio, J.M., Gonzalez, B.(2017) J Biological Chem 292: 10534-10548
- PubMed: 28450399 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M117.780395
- Primary Citation Related Structures:
5MW8, 5MWL, 5MWM - PubMed Abstract:
Inositol 1,3,4,5,6-pentakisphosphate 2-kinases (IP 5 2-Ks) are part of a family of enzymes in charge of synthesizing inositol hexakisphosphate (IP 6 ) in eukaryotic cells. This protein and its product IP 6 present many roles in cells, participating in mRNA export, embryonic development, and apoptosis. We reported previously that the full-length IP 5 2-K from Arabidopsis thaliana is a zinc metallo-enzyme, including two separated lobes (the N- and C-lobes). We have also shown conformational changes in IP 5 2-K and have identified the residues involved in substrate recognition and catalysis. However, the specific features of mammalian IP 5 2-Ks remain unknown. To this end, we report here the first structure for a murine IP 5 2-K in complex with ATP/IP 5 or IP 6 Our structural findings indicated that the general folding in N- and C-lobes is conserved with A. thaliana IP 5 2-K. A helical scaffold in the C-lobe constitutes the inositol phosphate-binding site, which, along with the participation of the N-lobe, endows high specificity to this protein. However, we also noted large structural differences between the orthologues from these two eukaryotic kingdoms. These differences include a novel zinc-binding site and regions unique to the mammalian IP 5 2-K, as an unexpected basic patch on the protein surface. In conclusion, our findings have uncovered distinct features of a mammalian IP 5 2-K and set the stage for investigations into protein-protein or protein-RNA interactions important for IP 5 2-K function and activity.
- From the Departamento de Cristalografía y Biología Estructural, Instituto de Química-Física "Rocasolano," Consejo Superior de Investigaciones Científicas, Serrano 119, 28006 Madrid, Spain and.
Organizational Affiliation: