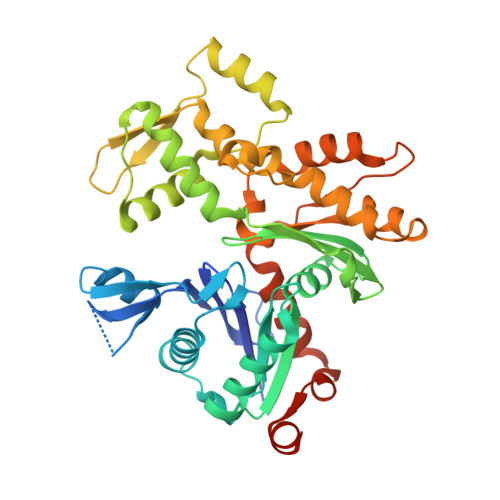
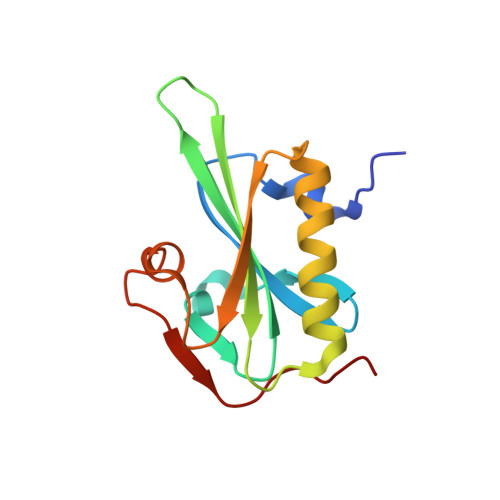
Rapid cadmium SAD phasing at the standard wavelength (1 angstrom ).
Panneerselvam, S., Kumpula, E.P., Kursula, I., Burkhardt, A., Meents, A.(2017) Acta Crystallogr D Struct Biol 73: 581-590
- PubMed: 28695858 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798317006970
- Primary Citation Related Structures:
5MVV, 5MYY - PubMed Abstract:
Cadmium ions can be effectively used to promote crystal growth and for experimental phasing. Here, the use of cadmium ions as a suitable anomalous scatterer at the standard wavelength of 1 Å is demonstrated. The structures of three different proteins were determined using cadmium single-wavelength anomalous dispersion (SAD) phasing. Owing to the strong anomalous signal, the structure of lysozyme could be automatically phased and built using a very low anomalous multiplicity (1.1) and low-completeness (77%) data set. Additionally, it is shown that cadmium ions can easily substitute divalent ions in ATP-divalent cation complexes. This property could be generally applied for phasing experiments of a wide range of nucleotide-binding proteins. Improvements in crystal growth and quality, good anomalous signal at standard wavelengths (i.e. no need to change photon energy) and rapid phasing and refinement using a single data set are benefits that should allow cadmium ions to be widely used for experimental phasing.
- Photon Science, DESY, Notkestrasse 85, 22607 Hamburg, Germany.
Organizational Affiliation: