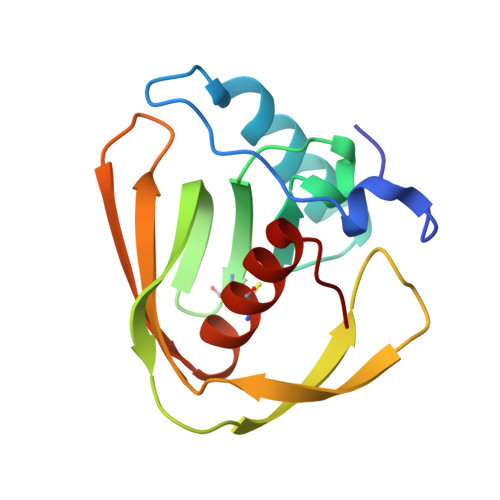
Peptide deformylases from Vibrio parahaemolyticus phage and bacteria display similar deformylase activity and inhibitor binding clefts.
Grzela, R., Nusbaum, J., Fieulaine, S., Lavecchia, F., Desmadril, M., Nhiri, N., Van Dorsselaer, A., Cianferani, S., Jacquet, E., Meinnel, T., Giglione, C.(2018) Biochim Biophys Acta 1866: 348-355
- PubMed: 29101077 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2017.10.007
- Primary Citation Related Structures:
5MTE - PubMed Abstract:
Unexpected peptide deformylase (PDF) genes were recently retrieved in numerous marine phage genomes. While various hypotheses dealing with the occurrence of these intriguing sequences have been made, no further characterization and functional studies have been described thus far. In this study, we characterize the bacteriophage Vp16 PDF enzyme, as representative member of the newly identified C-terminally truncated viral PDFs. We show here that conditions classically used for bacterial PDFs lead to an enzyme exhibiting weak activity. Nonetheless, our integrated biophysical and biochemical approaches reveal specific effects of pH and metals on Vp16 PDF stability and activity. A novel purification protocol taking in account these data allowed strong improvement of Vp16 PDF specific activity to values similar to those of bacterial PDFs. We next show that Vp16 PDF is as sensitive to the natural inhibitor compound of PDFs, actinonin, as bacterial PDFs. Comparison of the 3D structures of Vp16 and E. coli PDFs bound to actinonin also reveals that both PDFs display identical substrate binding mode. We conclude that bacteriophage Vp16 PDF protein has functional peptide deformylase activity and we suggest that encoded phage PDFs might be important for viral fitness.
- Institute for Integrative Biology of the Cell (I2BC), CEA, CNRS, Univ. Paris-Sud, Université Paris-Saclay, 91198 Gif-sur-Yvette Cedex, France.
Organizational Affiliation: