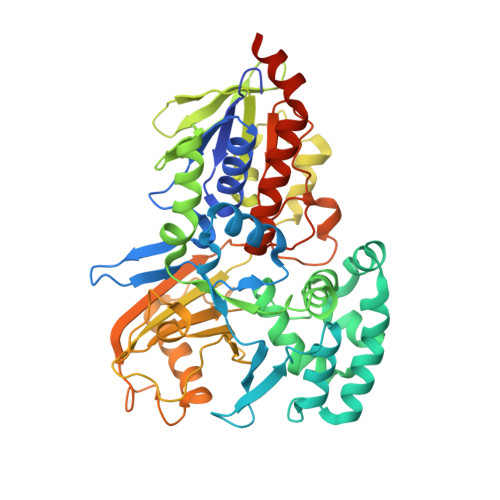
Structure of Phytoene Desaturase Provides Insights into Herbicide Binding and Reaction Mechanisms Involved in Carotene Desaturation.
Brausemann, A., Gemmecker, S., Koschmieder, J., Ghisla, S., Beyer, P., Einsle, O.(2017) Structure 25: 1222-1232.e3
- PubMed: 28669634 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2017.06.002
- Primary Citation Related Structures:
5MOG - PubMed Abstract:
Cyanobacteria and plants synthesize carotenoids via a poly-cis pathway starting with phytoene, a membrane-bound C40 hydrocarbon. Phytoene desaturase (PDS) introduces two double bonds and concomitantly isomerizes two neighboring double bonds from trans to cis. PDS assembles into homo-tetramers that interact monotopically with membranes. A long hydrophobic tunnel is proposed to function in the sequential binding of phytoene and the electron acceptor plastoquinone. The herbicidal inhibitor norflurazon binds at a plastoquinone site thereby blocking reoxidation of FAD red . Comparison with the sequence-dissimilar bacterial carotene desaturase CRTI reveals substantial similarities in the overall protein fold, defining both as members of the GR2 family of flavoproteins. However, the PDS active center architecture is unprecedented: no functional groups are found in the immediate flavin vicinity that might participate in dehydrogenation and isomerization. This suggests that the isoalloxazine moiety is sufficient for catalysis. Despite mechanistic differences, an ancient evolutionary relation of PDS and CRTI is apparent.
- Institute for Biochemistry, Albert-Ludwigs-Universität Freiburg, Albertstrasse 21, 79104 Freiburg, Germany.
Organizational Affiliation: