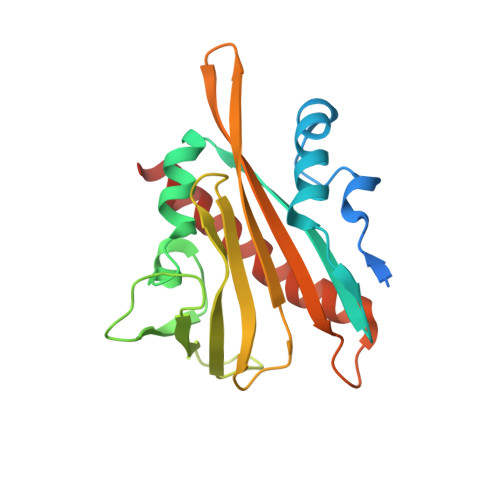
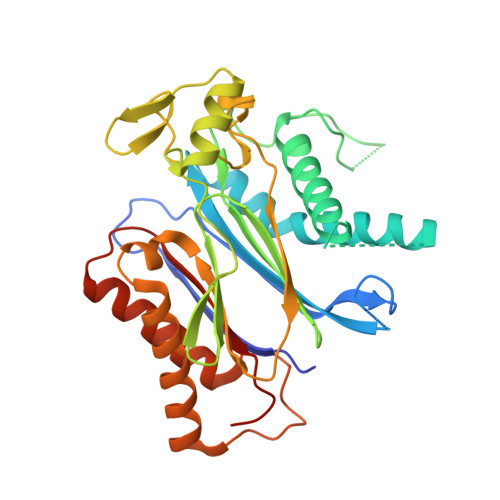
Structure of Ligand-Bound Intermediates of Crop ABA Receptors Highlights PP2C as Necessary ABA Co-receptor.
Moreno-Alvero, M., Yunta, C., Gonzalez-Guzman, M., Lozano-Juste, J., Benavente, J.L., Arbona, V., Menendez, M., Martinez-Ripoll, M., Infantes, L., Gomez-Cadenas, A., Rodriguez, P.L., Albert, A.(2017) Mol Plant 10: 1250-1253
- PubMed: 28736053 Search on PubMed
- DOI: https://doi.org/10.1016/j.molp.2017.07.004
- Primary Citation Related Structures:
5MMQ, 5MMX, 5MN0, 5MOA, 5MOB - Instituto de Química Física Rocasolano, Consejo Superior de Investigaciones Científicas, 28006 Madrid, Spain.
Organizational Affiliation: